What is KnockTF 2.0?
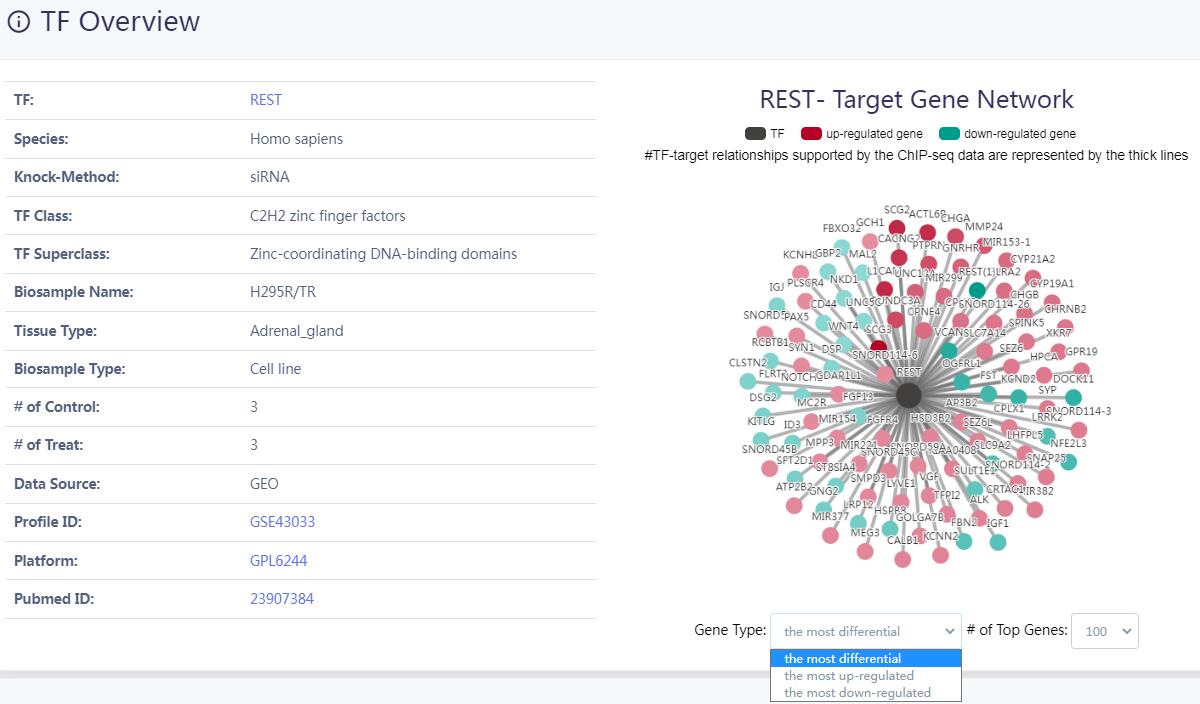
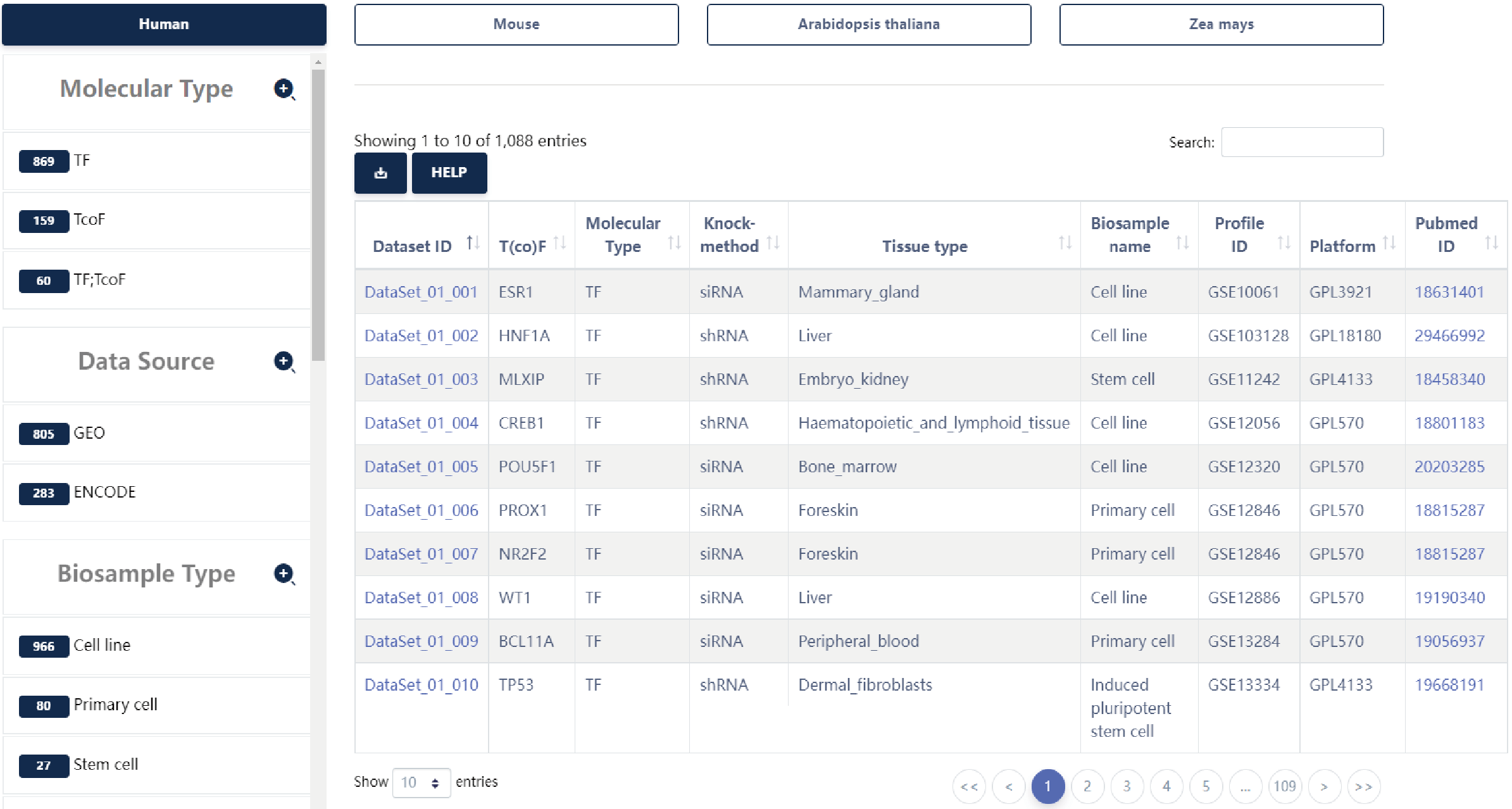
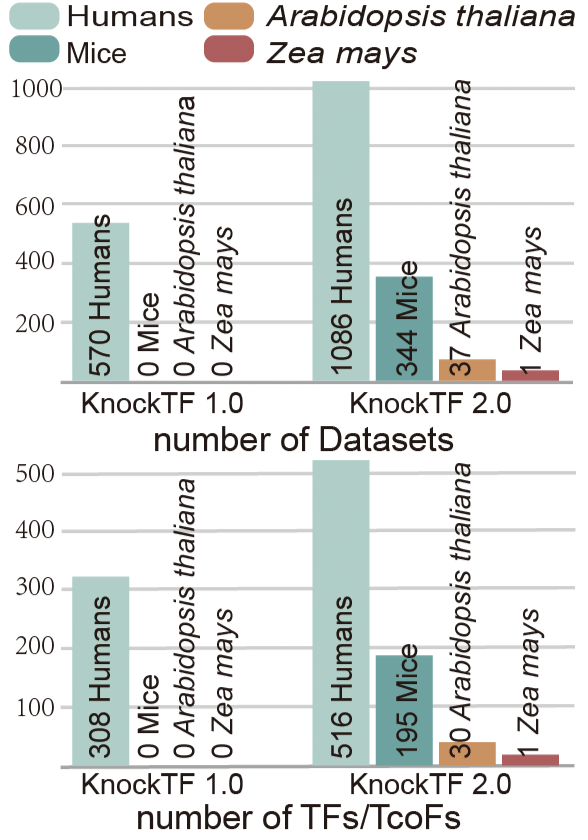
KnockTF 2.0 aims to provide comprehensive T(co)F knockdown/knockout dataset resource across multiple tissue/cell types of different species. The current version of KnockTF stores 1468 manually curated RNA-seq and microarray datasets associated with 612 TFs and 172 TcoFs disrupted by different knockdown/knockout techniques and across multiple tissue/cell types in humans, mice, Arabidopsis thaliana and Zea mays.
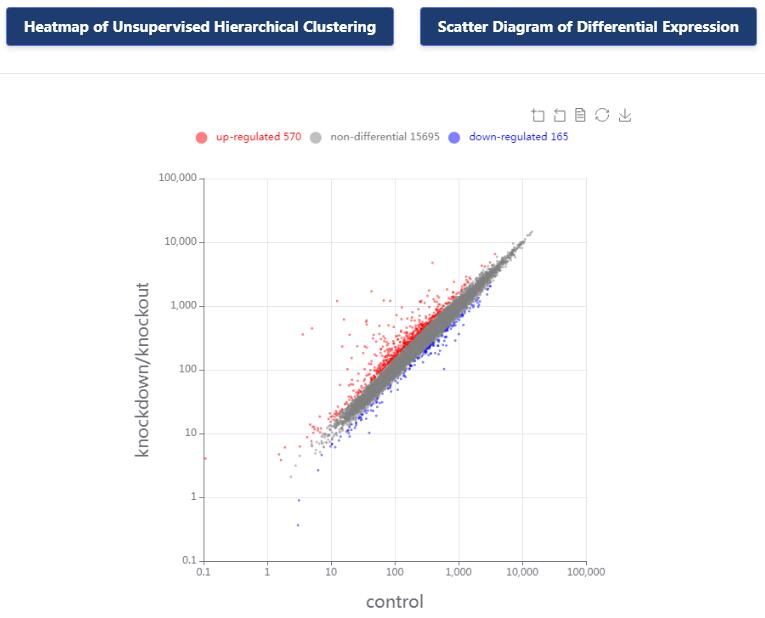
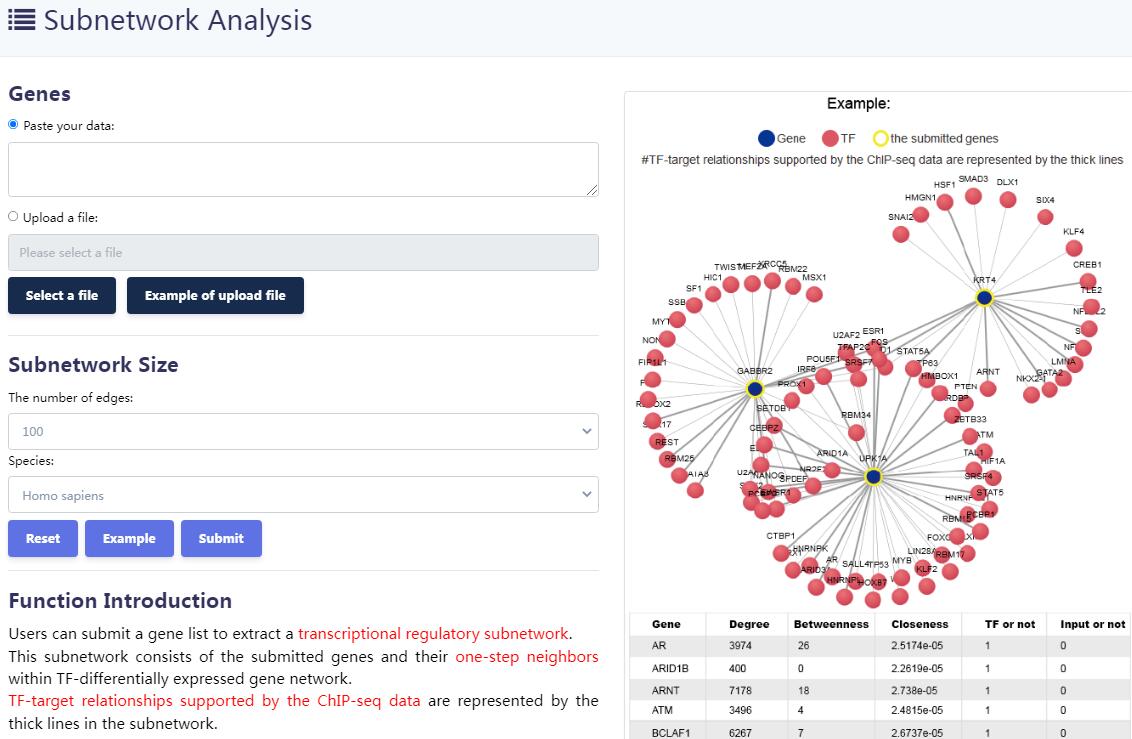
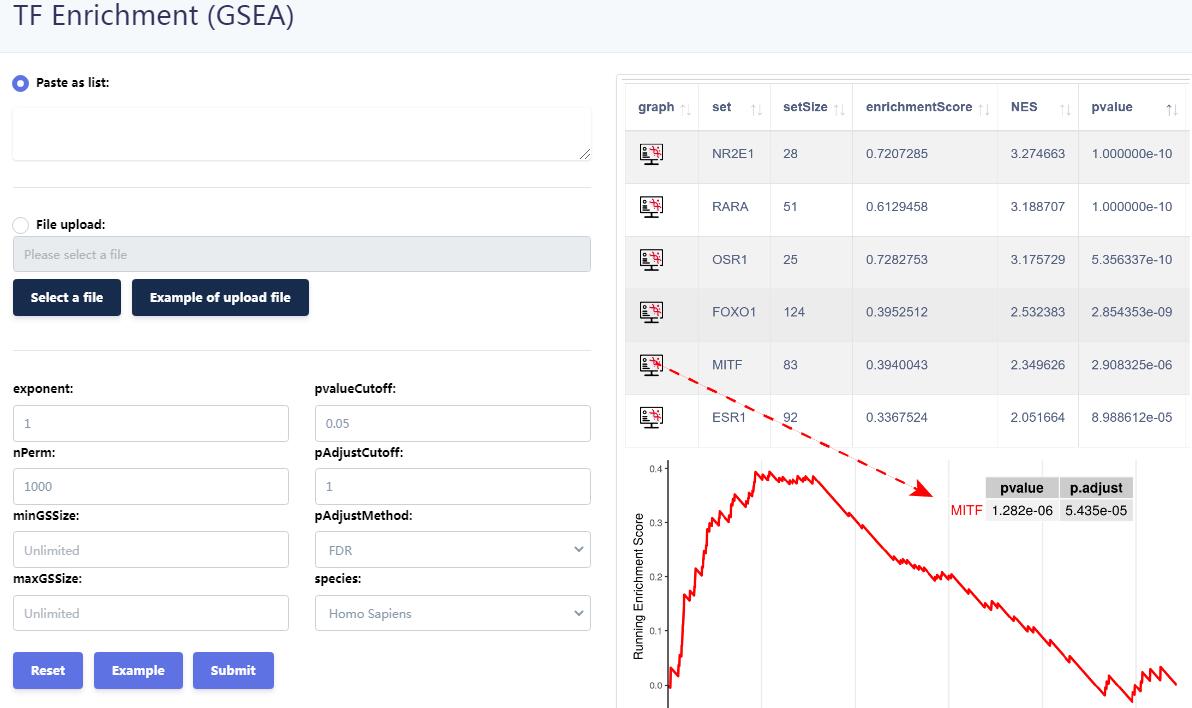
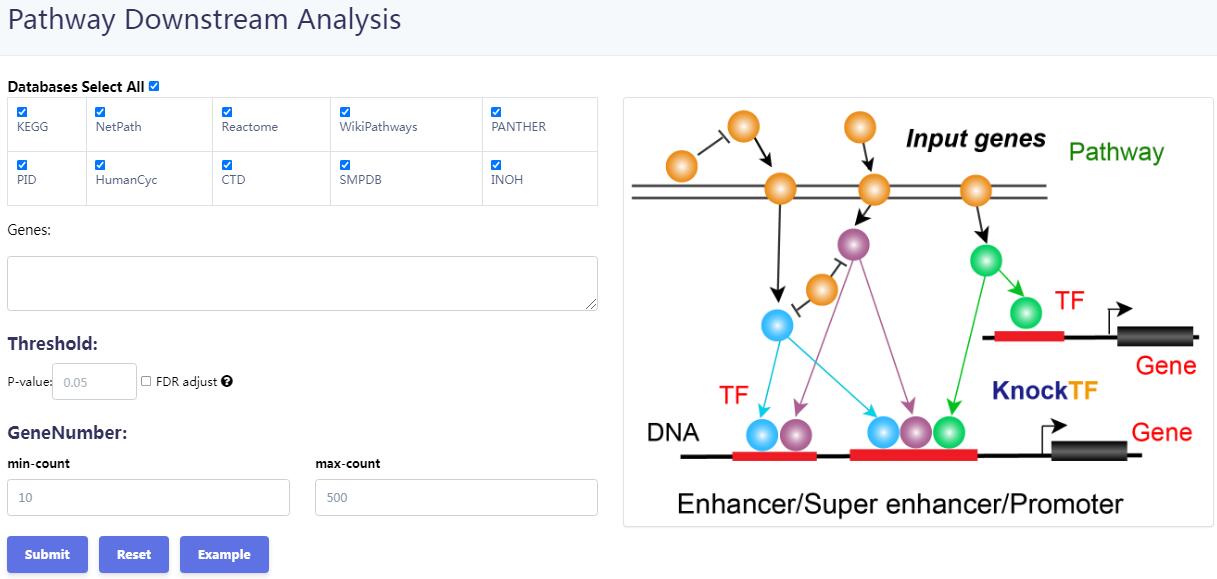
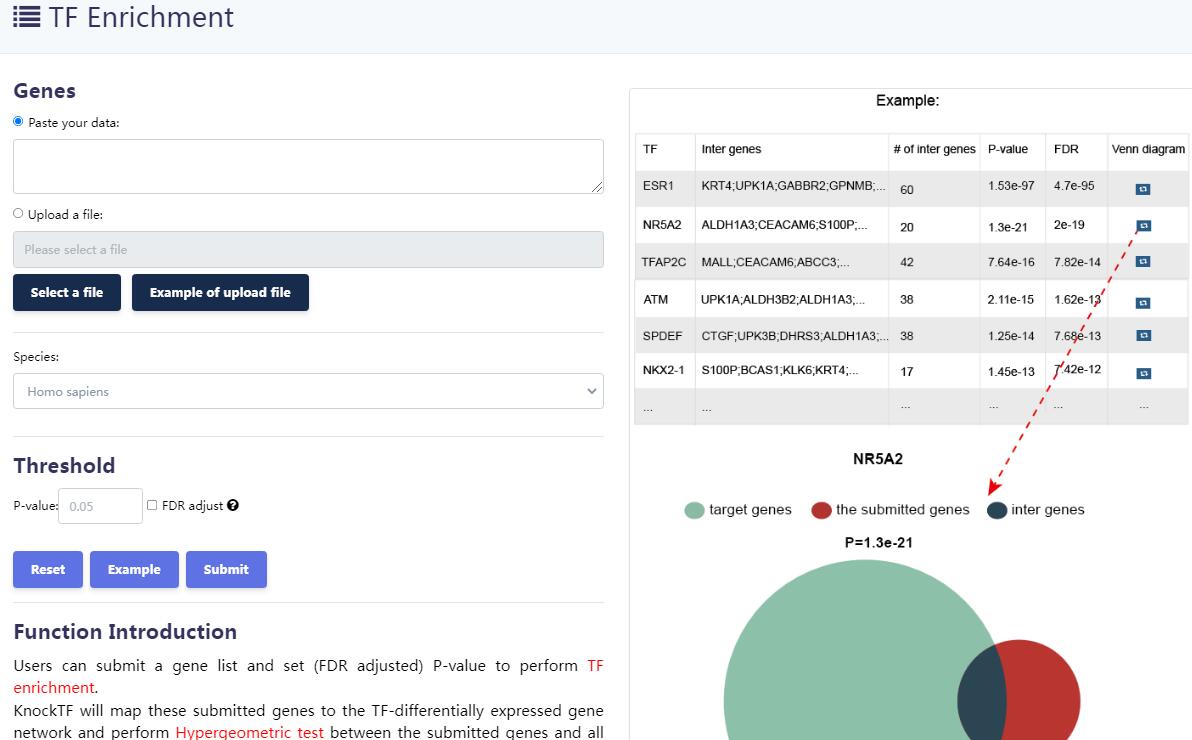
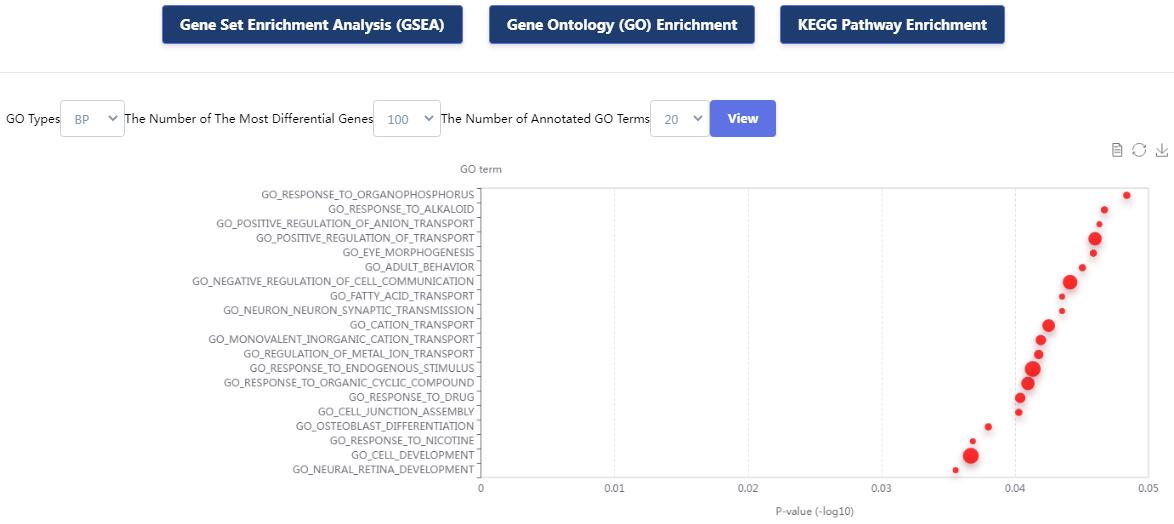
KnockTF 2.0 not only provides comprehensive gene expression information for T(co)F target genes of interest, but also collects upstream pathway information for T(co)Fs and various functional annotation information for downstream target genes, such as GO/KEGG pathway enrichment, hierarchical clustering analysis and differentially expressed analysis. Furthermore, KnockTF 2.0 provides the detailed and abundant (epi)genetic annotation information for T(co)F target genes, including super-enhancers, enhancers, transcription factor binding sites (TFBSs), common SNPs, risk SNPs, LD SNPs, expression quantitative trait locus (eQTL), methylation sites, DNase I hypersensitivity sites (DHSs), chromatin interactions, chromatin accessibility regions, CRISPR/Cas9 target sites, and topologically associating domains (TADs).
KnockTF 2.0 will facilitate the identification of functional T(co)Fs and target genes, and benefit the investigation of their roles in the physiological and pathological processes.

TF-Marker
TF-Marker database for human
TcoFbase
Document a large number of available resources of mammals TcoFs
SEdb
SEdb: The comprehensive human Super-Enhancer databasebr
SEanalysis
SEanalysis: a web tool for super-enhancer associated regulatory analysis
ENdb
ENdb: An experimentally supported enhancer database for human and mouse
VARAdb
VARAdb: A comprehensive human variation annotation database
TRlnc
TRlnc: A comprehensive database of human transcriptional regulation of lncRNAs
TRCirc
TRCirc: A resource for transcriptional regulation information of circRNAs
ATACdb
ATACdb: A comprehensive human chromatin accessibility database
LncSEA
LncSEA: A comprehensive human lncRNA sets resource and enrichment analysis platform
Principal Investigator: Chunquan Li, Ph.D.
The First Affiliated Hospital, Hengyang Medical School, University of South China
Email: lcqbio@163.com
We welcome researchers from all over the world to provide valuable advice for KnockTF, and make KnockTF more and more perfect.
Feng C, Song C, Liu Y, Qian F, Gao Y, Ning Z, Wang Q, Jiang Y, Li Y, Li M, Chen J, Zhang J, Li C. KnockTF: a comprehensive human gene expression profile database with knockdown/knockout of transcription factors. Nucleic Acids Res. 2020 Jan 8;48(D1):D93-D100. doi: 10.1093/nar/gkz881. PMID: 31598675; PMCID: PMC6943067.