Welcome to TcoFBase !
What is TcoFBase?
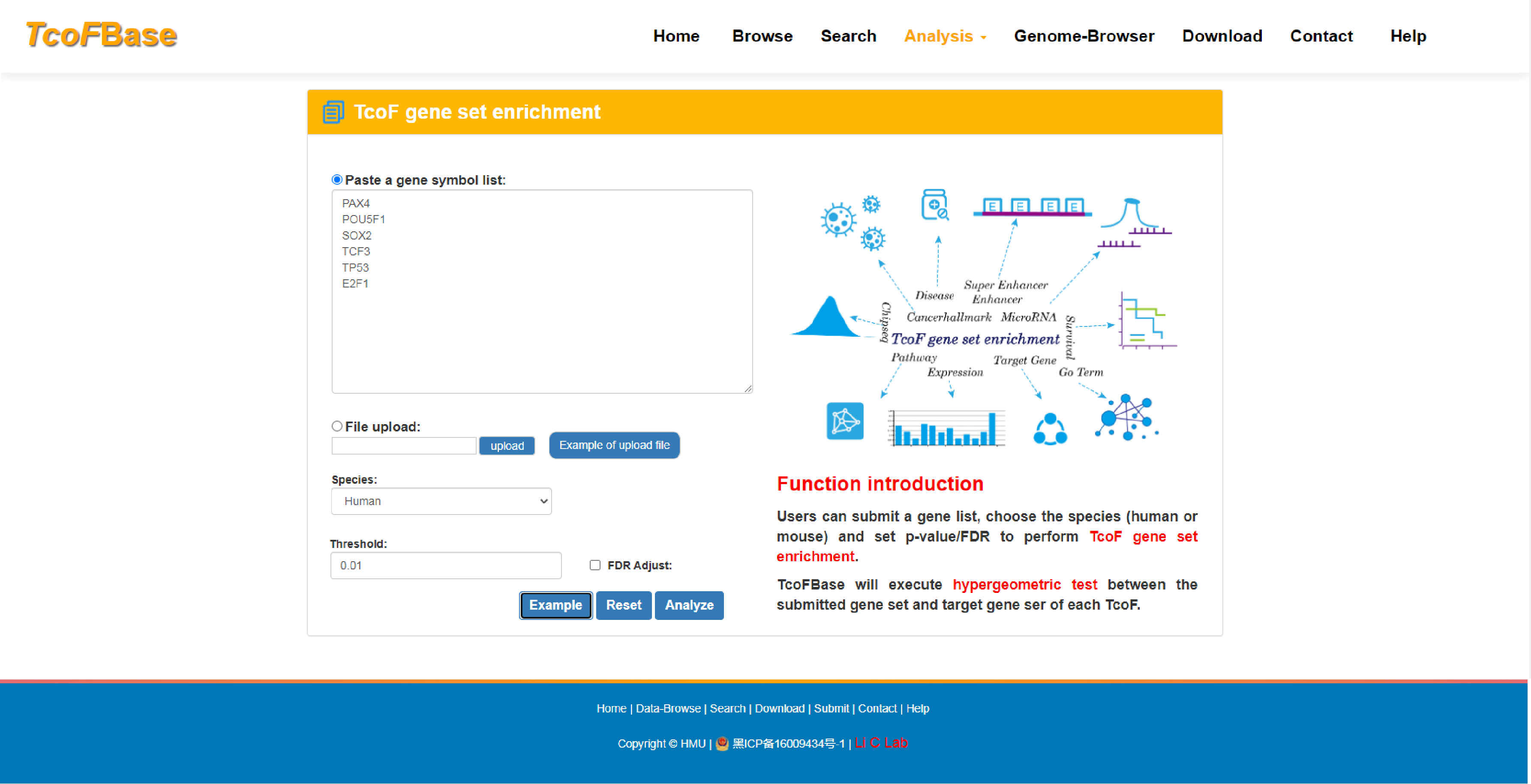
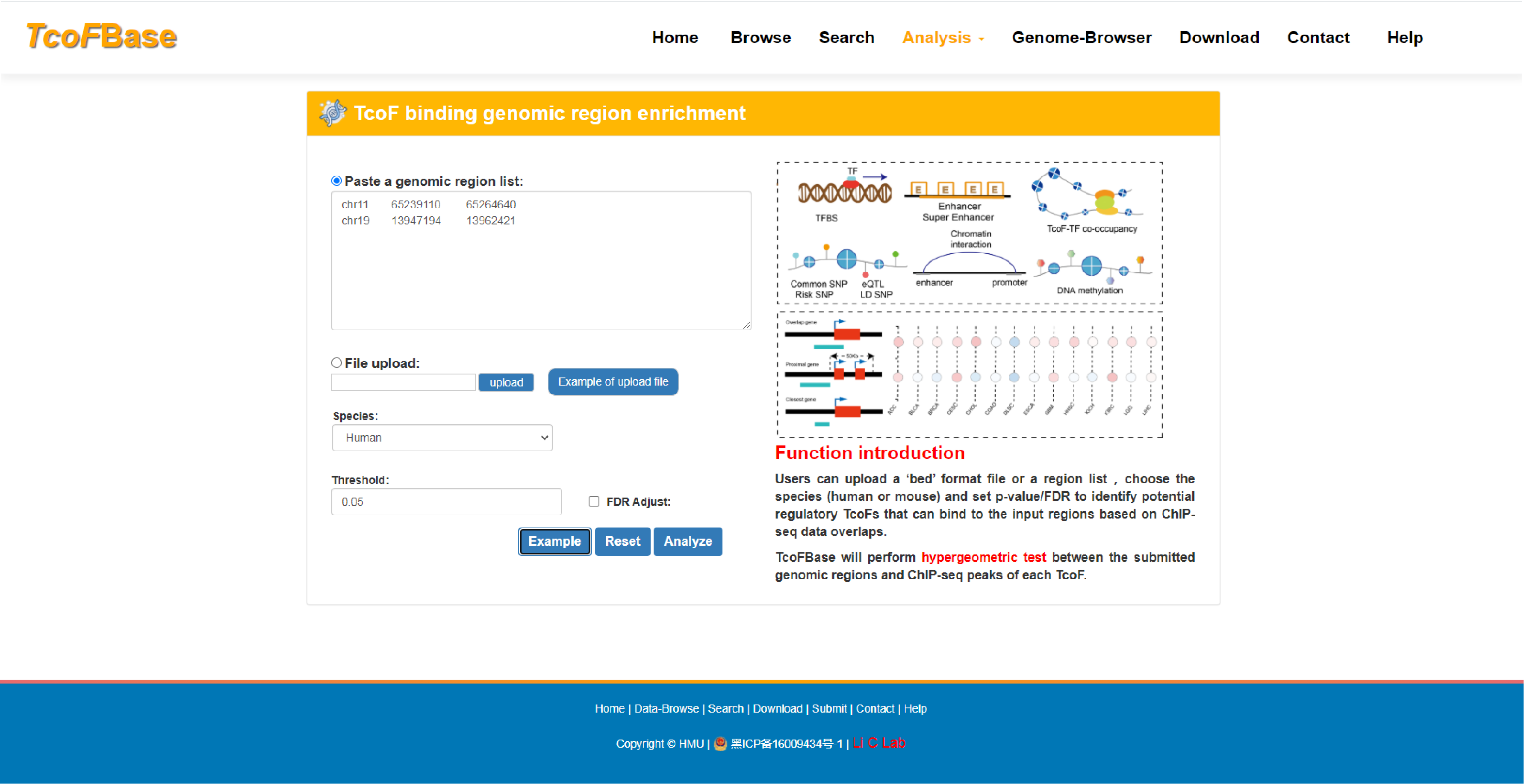
Here, we developed the TcoFBase database, which aims to document a large number of available resources of mammals TcoFs, provided annotation and enrichment analyses of TcoFs.
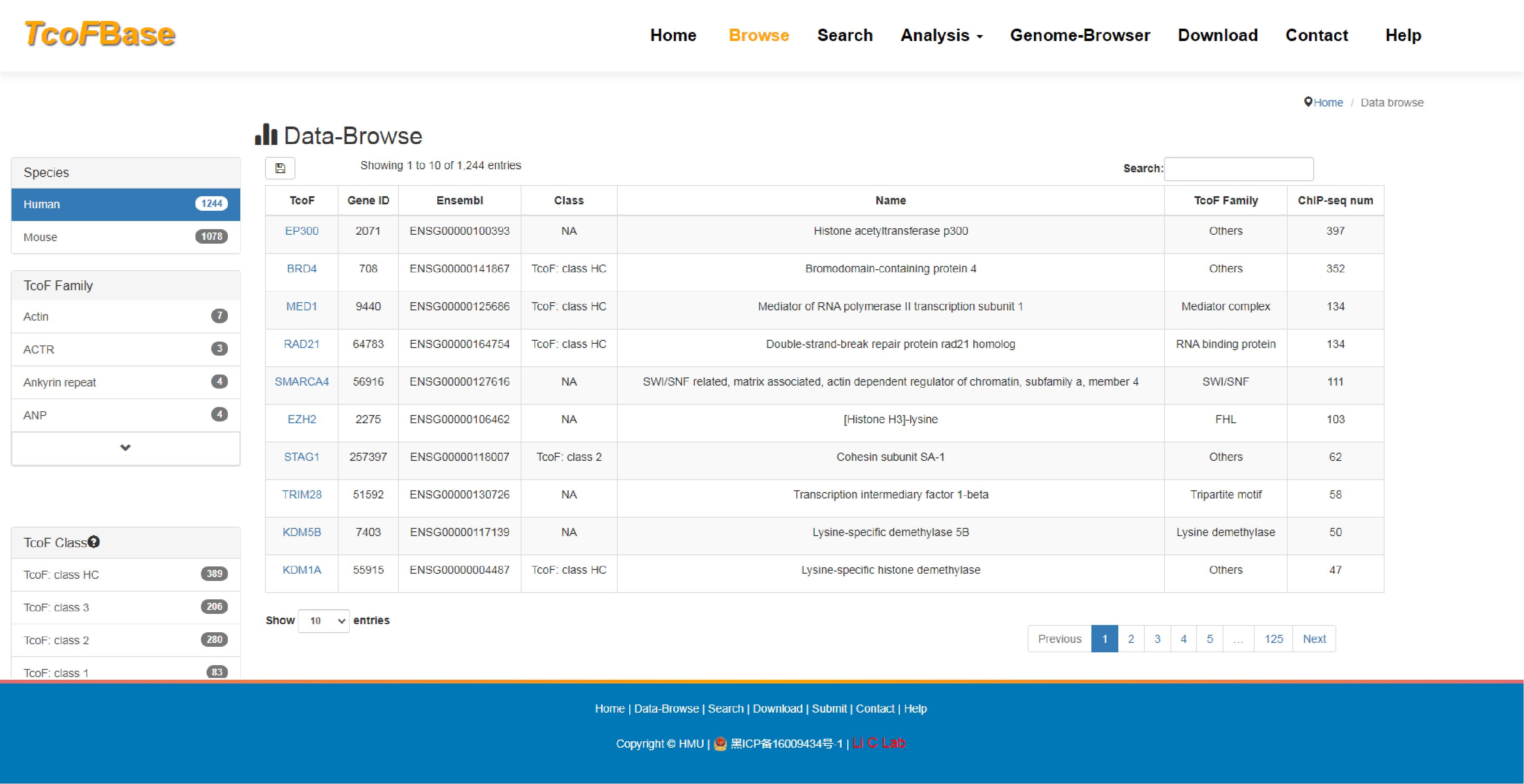
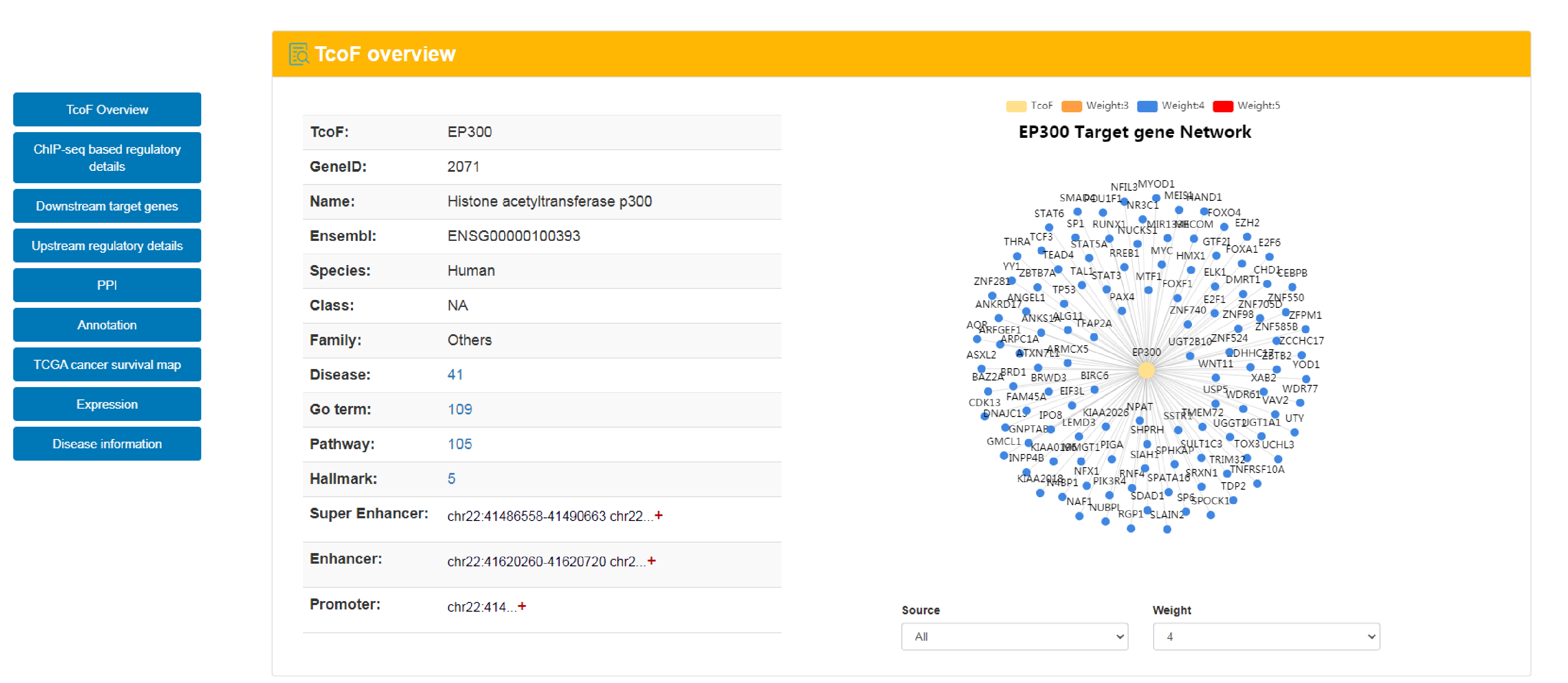
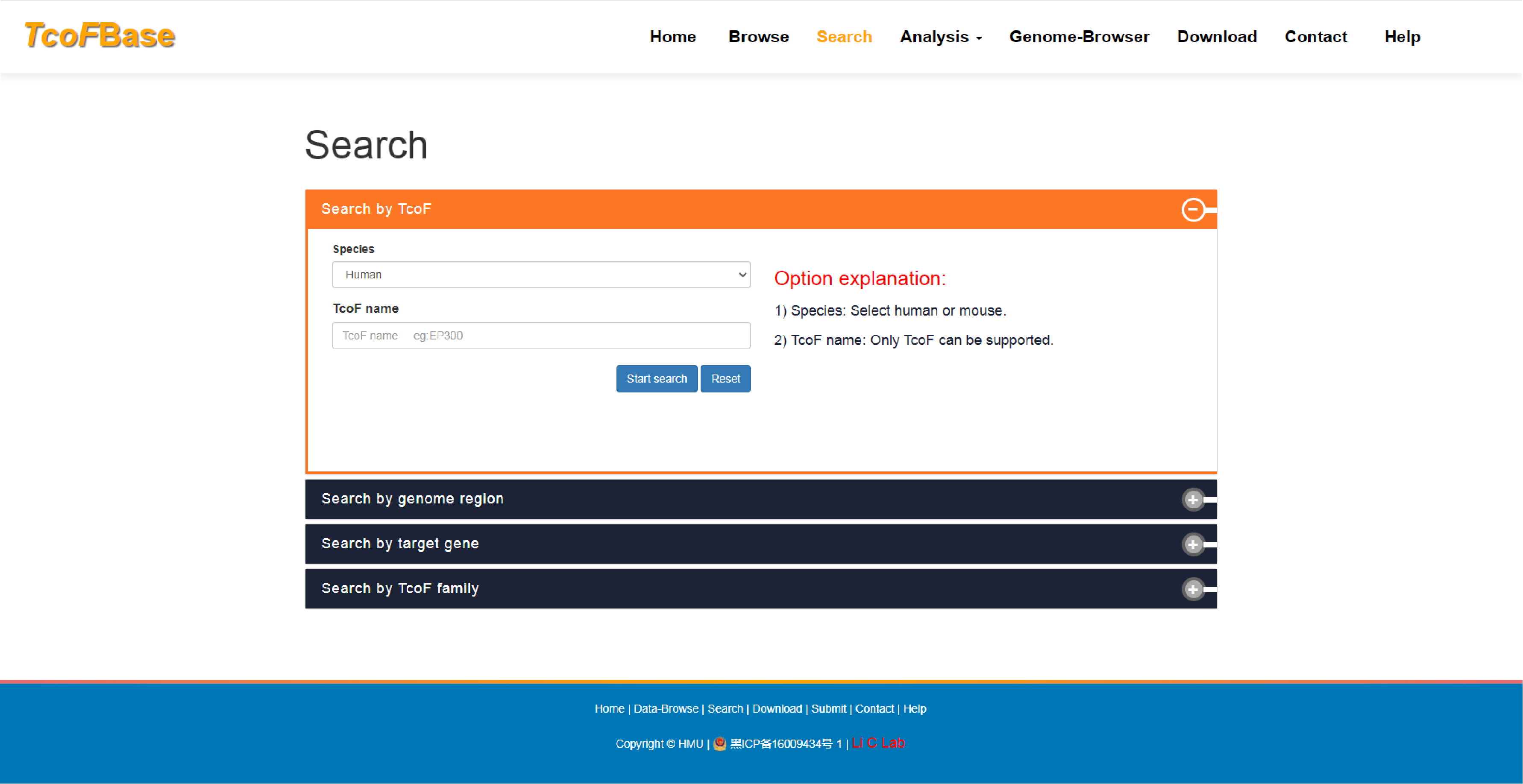
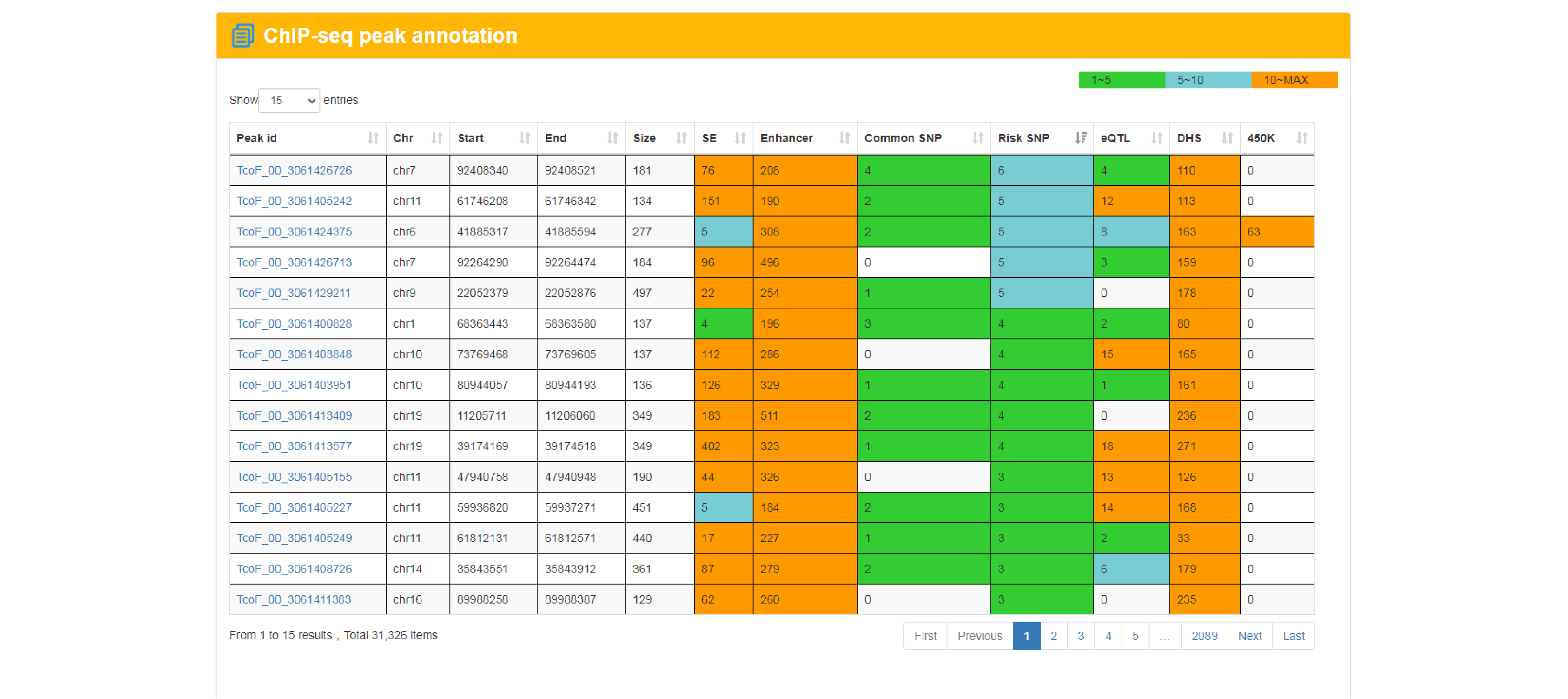
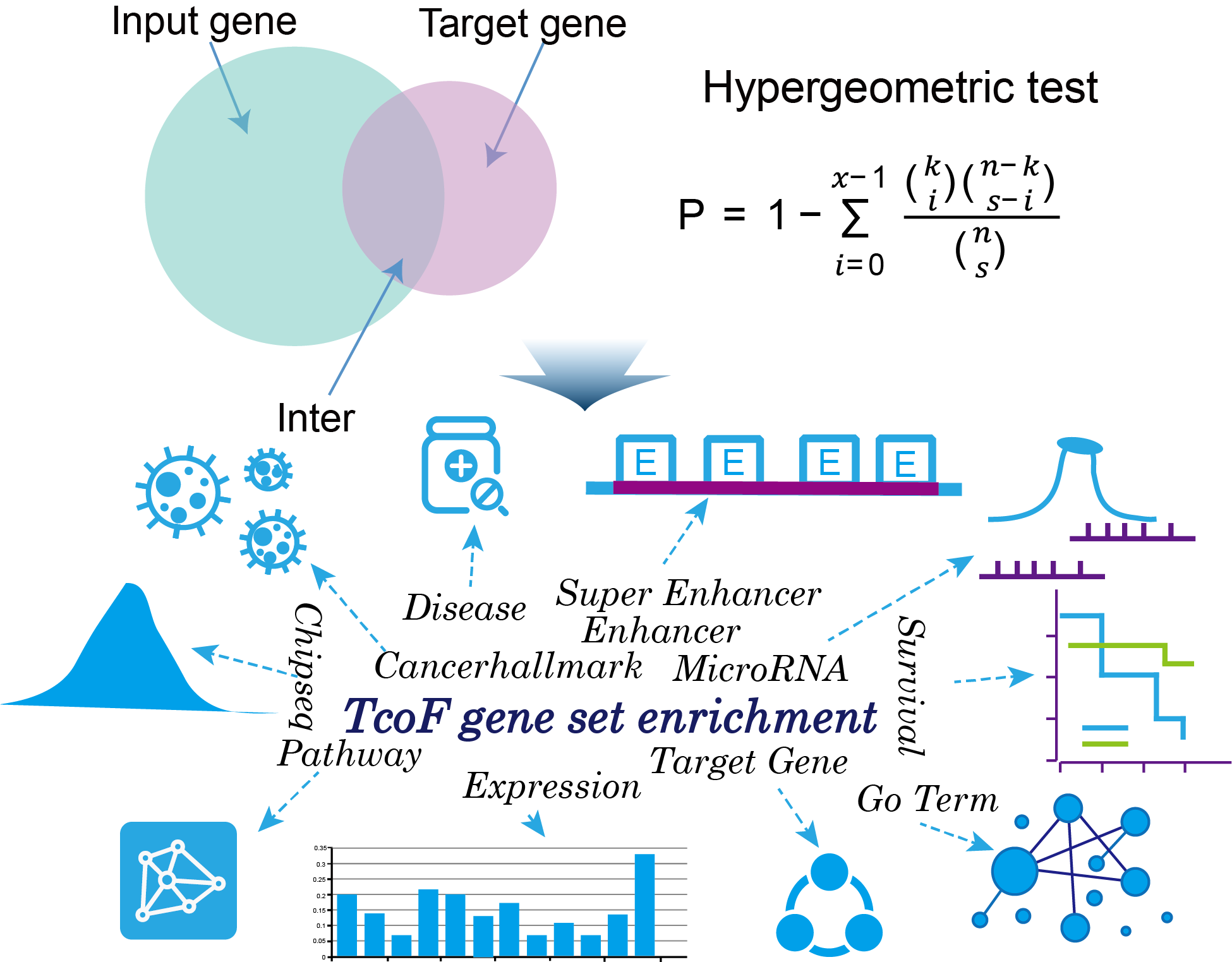
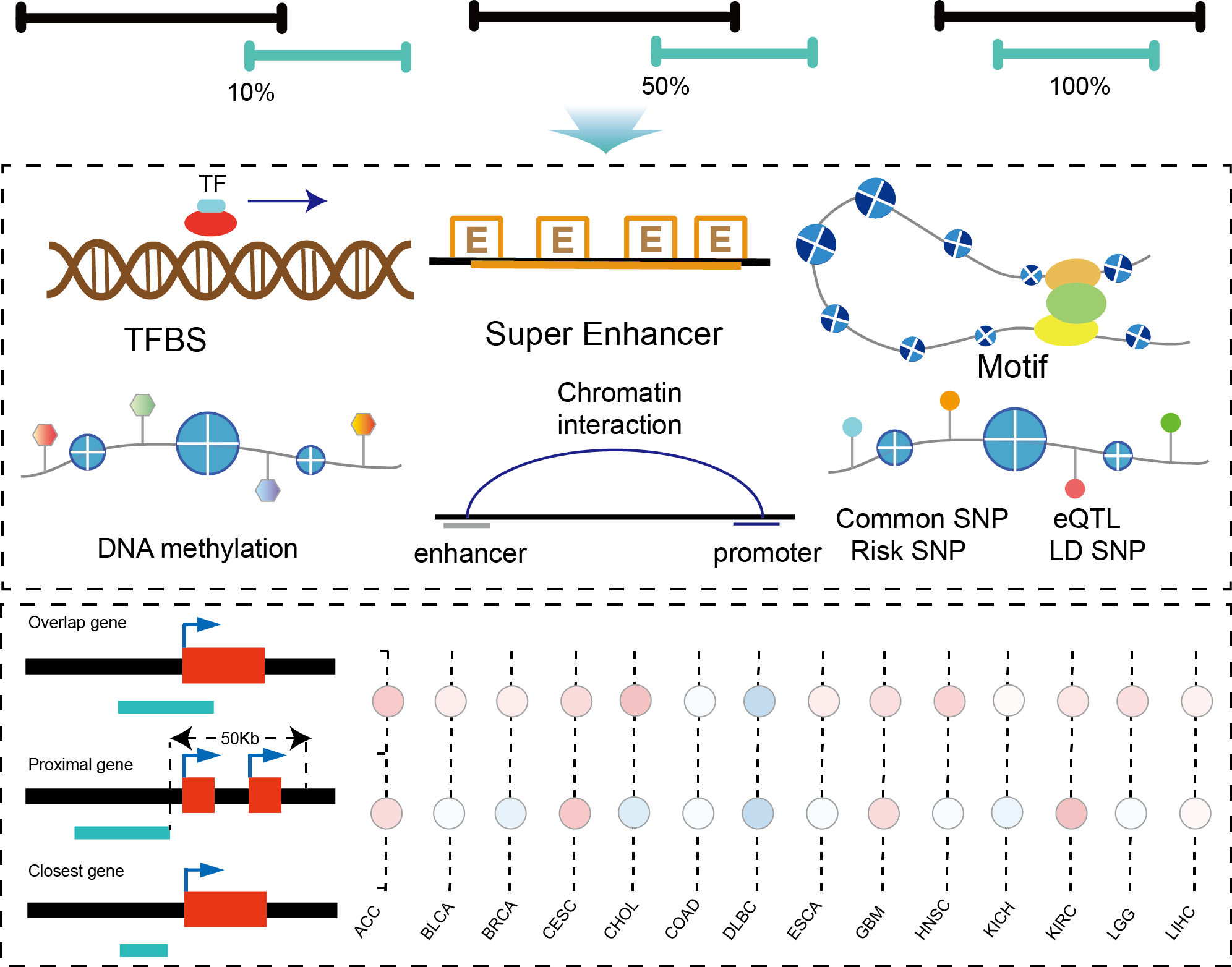
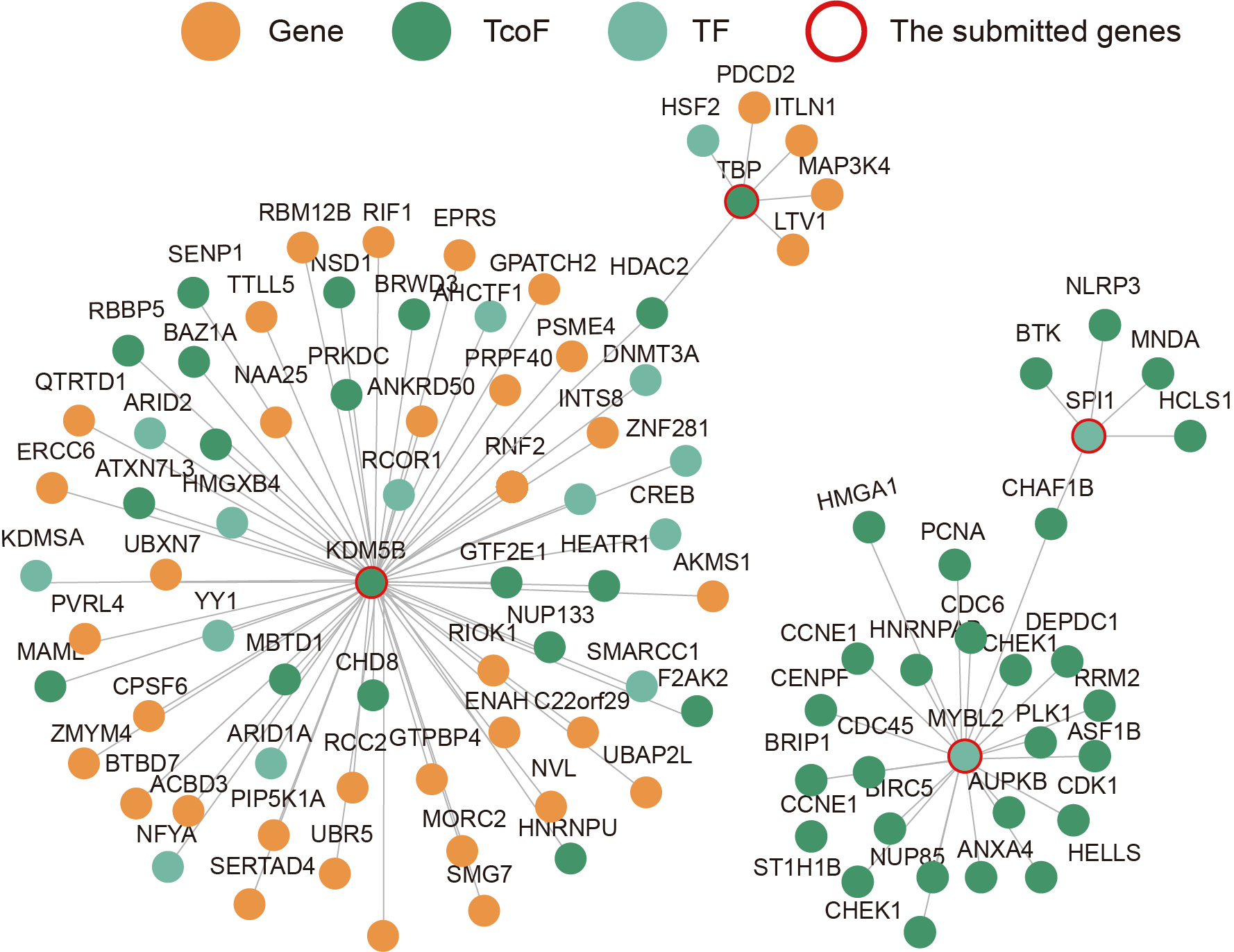
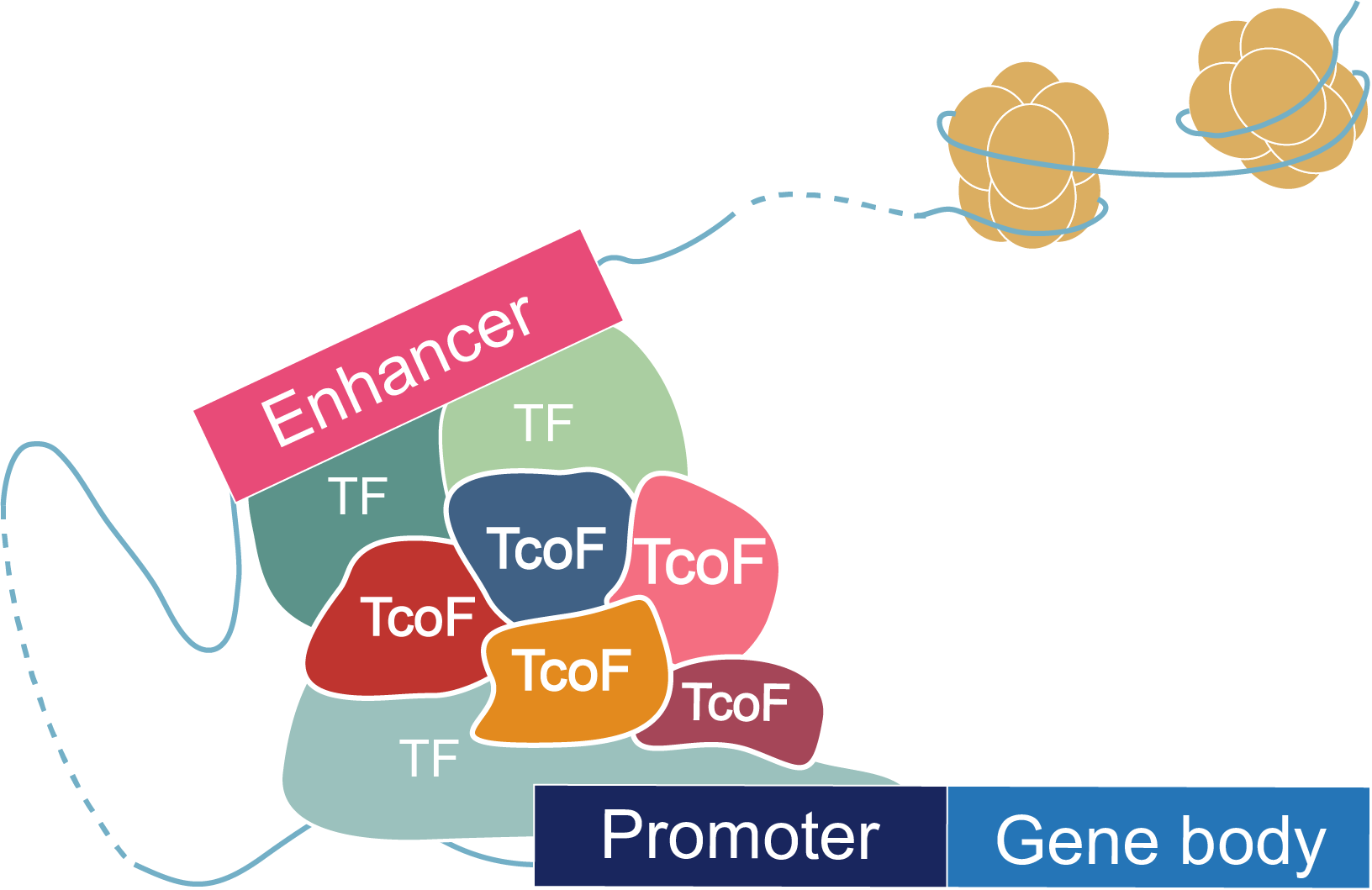
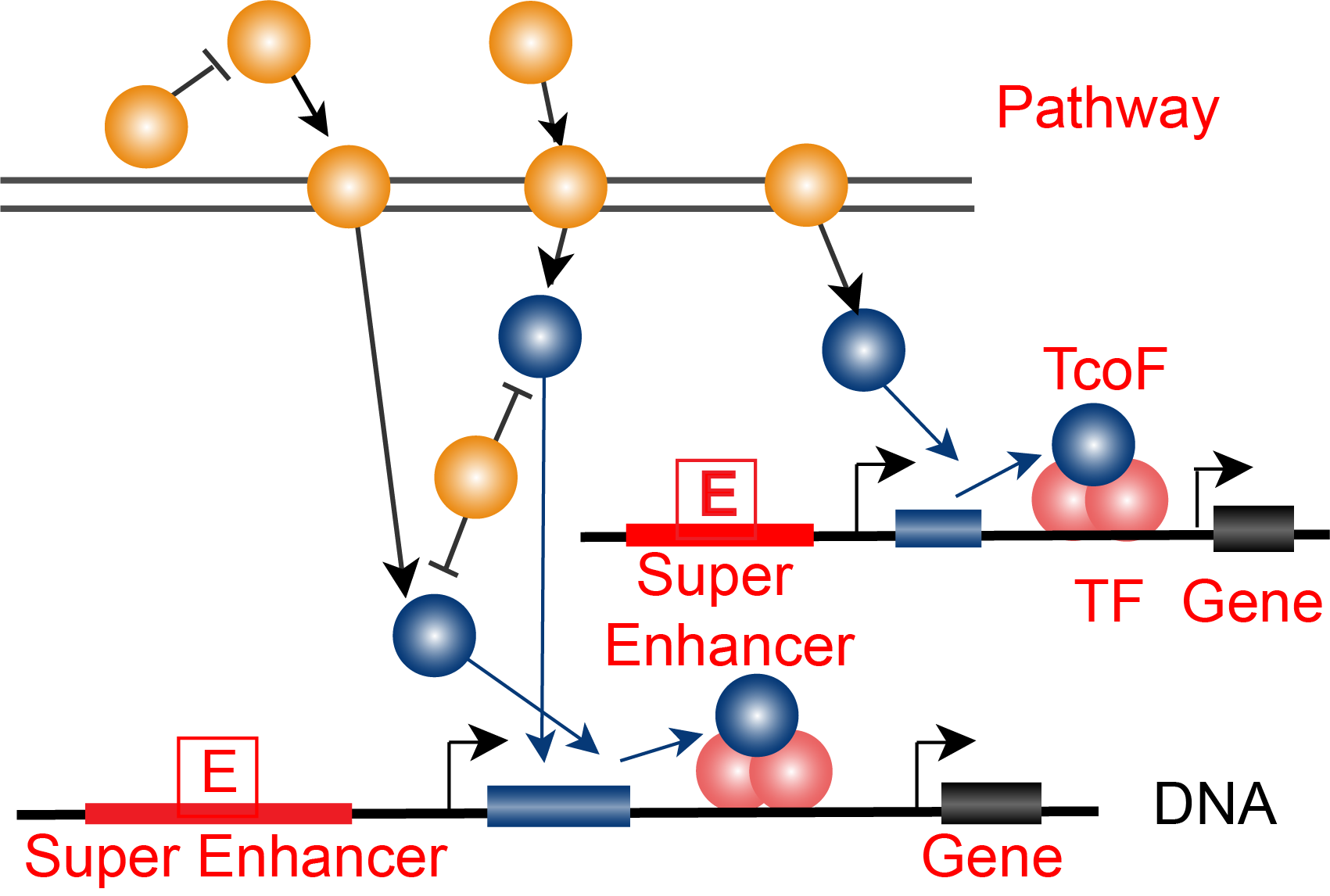
The current version of TcoFBase cataloged a total of 2,322 TcoFs and 5,673 TcoFs associated ChIP-seq data from over 380 tissues/cell types in human and mouse. Emphatically, TcoFBase provides the detailed and abundant (epi) genetic annotations in ChIP-seq based TcoF binding regions, such as super enhancer, enhancer, TFBS, methylation sites, common SNPs, risk SNPs, eQTLs, histone modifications and 3D chromatin interactions. TcoFBase also provides TcoFs downstream target genes by mapping binding regions into genomes in five methods. Furthermore, TcoFBase provides upstream regulatory information for TcoFs and various functional annotation including pathway, GO Trem, cancer hallmark, PPI, survival, expression and disease information. Especially, TcoFBase provides 5 types of TcoF regulatory analyses for users, including TcoF gene set enrichment, TcoF binding genomic region annotation, TcoF regulatory network analysis, TcoF-TF co-occupancy analysis, TcoF regulatory axis analysis. TcoFBase is a user-friendly database to query, browse and visualize information associated with TcoF. We believe that TcoFBase could become a useful and effective platform for exploring potential functions and regulation of TcoF in diseases and biological processes.
TcoFs-associated Analysis
Data Statistical Table
| TcoF | The Number of Sources | 2 |
| The Number of TcoFs | 2322 | |
| TcoF(ChIP-seq) | The Number of Sources | 4 |
| The Number of TcoFs | 497 | |
| The Number of Samples | 6759 | |
| TF(Motif) | The Number of Sources | 6 |
| The Number of TFs | 699 | |
| The Number of Motifs | 3279 | |
| Pathway | The Number of Sources | 10 |
| The Number of Pathways | 2169 |
Sister Projects
ENdb
ENdb: an experimentally supported enhancer database for human and mouse
SEanalysis
SEanalysis: a web tool for super-enhancer associated regulatory analysis
VARAdb
VARAdb: a variation annotation database for human
ATACdb
ATACdb: a comprehensive human chromatin accessibility database
SEdb
SEdb: the comprehensive human Super-Enhancer database
KnockTF
KnockTF: a comprehensive human gene expression profile database with knockdown/knockout of transcription
factors
TRCirc
TRCirc: a resource for transcriptional regulation information of circRNAs
TRlnc
TRlnc: a comprehensive database of human transcriptional regulation of lncRNAs
LncSEA
LncSEA: a comprehensive human lncRNA sets resource and enrichment analysis platform
Contact us
Principal Investigator: Chunquan Li, Ph.D.
School of Medical Informatics,Daqing Campus Harbin Medical University 39 Xinyang Road, Daqing 163319, China
Phone: 86-459-8153035
Fax: 86-459-8153035
Email: lcqbio@163.com
Publication
Zhang Y, Song C, Zhang Y, Wang Y, Feng C, Chen J, Wei L, Pan Q, Shang D, Zhu Y, Zhu J, Fang S, Zhao J, Yang Y, Zhao X, Xu X, Wang Q, Guo J, Li C. TcoFBase: a comprehensive database for decoding the regulatory transcription co-factors in human and mouse. Nucleic Acids Res. 2022 Jan 7;50(D1):D391-D401. doi:10.1093/nar/gkab950 PMID: 34718747; PMCID: PMC8728270.