1. What information is available in the TF-Marker?
Transcription factors (TFs) recognize specific DNA sequences to control chromatin and transcription, forming a complex system that guides expression of the genome. Individual cells represent the basic blocks of tissues and organisms. Markers are signatures that help researchers to identify cell and tissue types. Many studies have confirmed that the regulation between TFs and related markers can drive cell differentiation and influence human diseases and biological processes. TFs coordinate cell type-specific transcriptional programs typically. Core transcriptional regulatory circuit (CRC) is comprised of a group of interconnected auto-regulating TFs forming loops, and core TFs in CRCs have been proved highly valuable for cell-type-specific transcriptional regulation in healthy and disease cells.Here, we established a comprehensive manually curated TFs, markers database for human (TF-Marker, http://bio.liclab.net/TF-Marker/index.php). Through reviewing 2091 published literatures, the current release of TF-Marker documents 1,316 TFs, 1,092 T Marker, 473 I Marker , 1,600 TFMarker, 1,424 TF Pmarker, involving 383 cell types and 95 tissue types for human. TF-Marker will help identify specific cell types with signature genes such as transcription factors,markers and TFMarkers in analyzing sing-cell Sequencing data.
2. Database content and construction
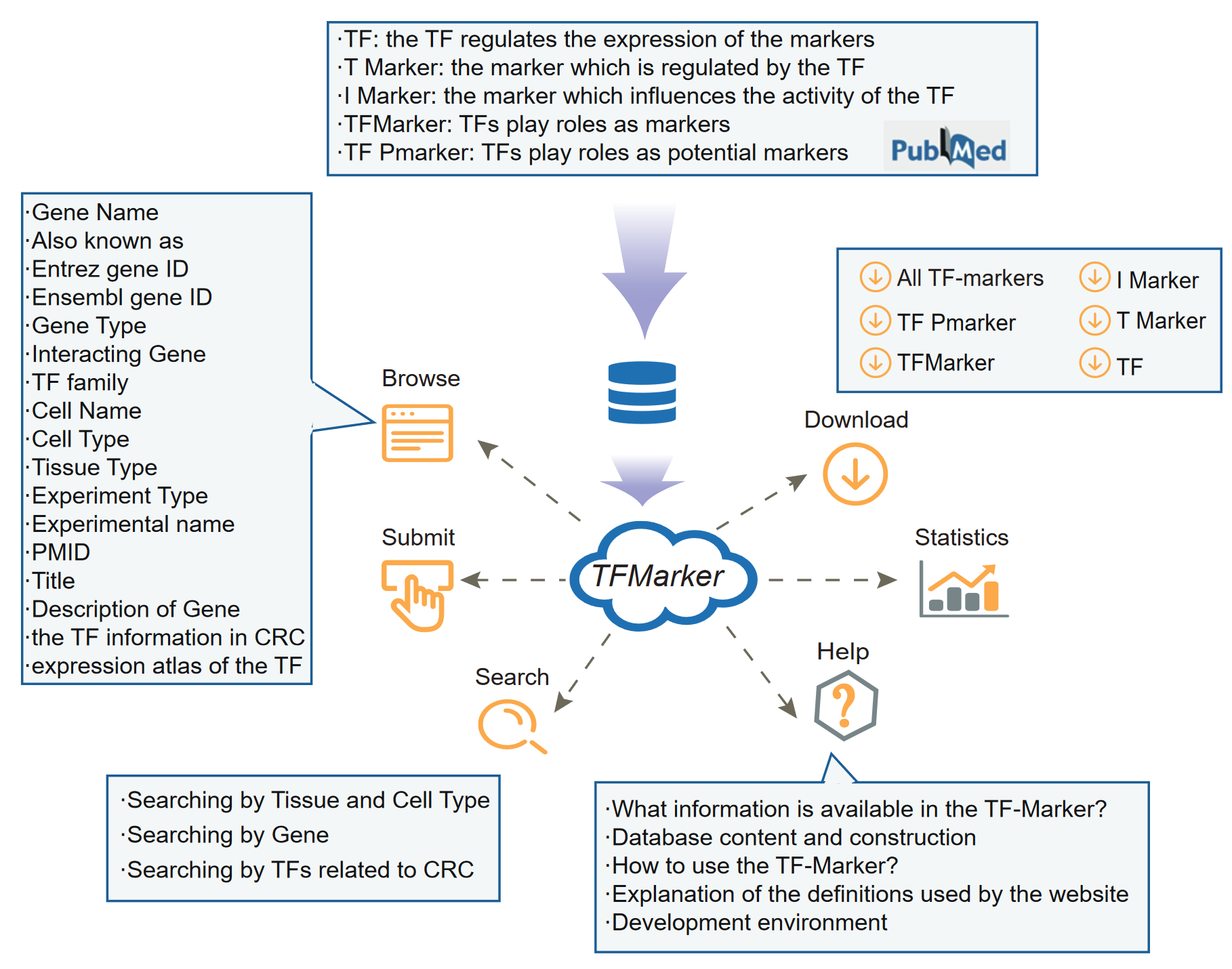
Through reviewing 2091 published literature, we considered the genes manually curated to five types, according to their functions: 1) TFs: TFs regulate the expression of the markers; 2) T Marker: it is the marker which is regulated by the TFs; 3) I Marker: it is the marker which influences the activity of the TFs; 4) TFMarker: TFs play roles as markers;5) TF Pmarker: TFs play roles as potential markers. We manually examined TFs and related markers in each published paper, including 1,316 TFs, 1,092 T Marker, 473 I Marker , 1,600 TFMarker, 1,424 TF Pmarker, involving 383 cell types and 95 tissue types for human. We further collected the detailed information of these TFs and related markers, which were supported by strong experimental evidence including RNAi, in vitro knockdown, western blot, qRT-PCR, luciferase reporter assay and Immunofluorescent staining. TF-Marker provides a user-friendly interface to browse, query, and visualize the detailed information about the TFs and related markers.
3. How to use the TF-Marker?
3.1. Search
Users can search genes through four ways, including 'Search by tissue and cell type', 'Search by gene name' and 'Search by TFs related to Core transcriptional Regulatory Circuit'.

3.1.1. Search by tissue and cell type
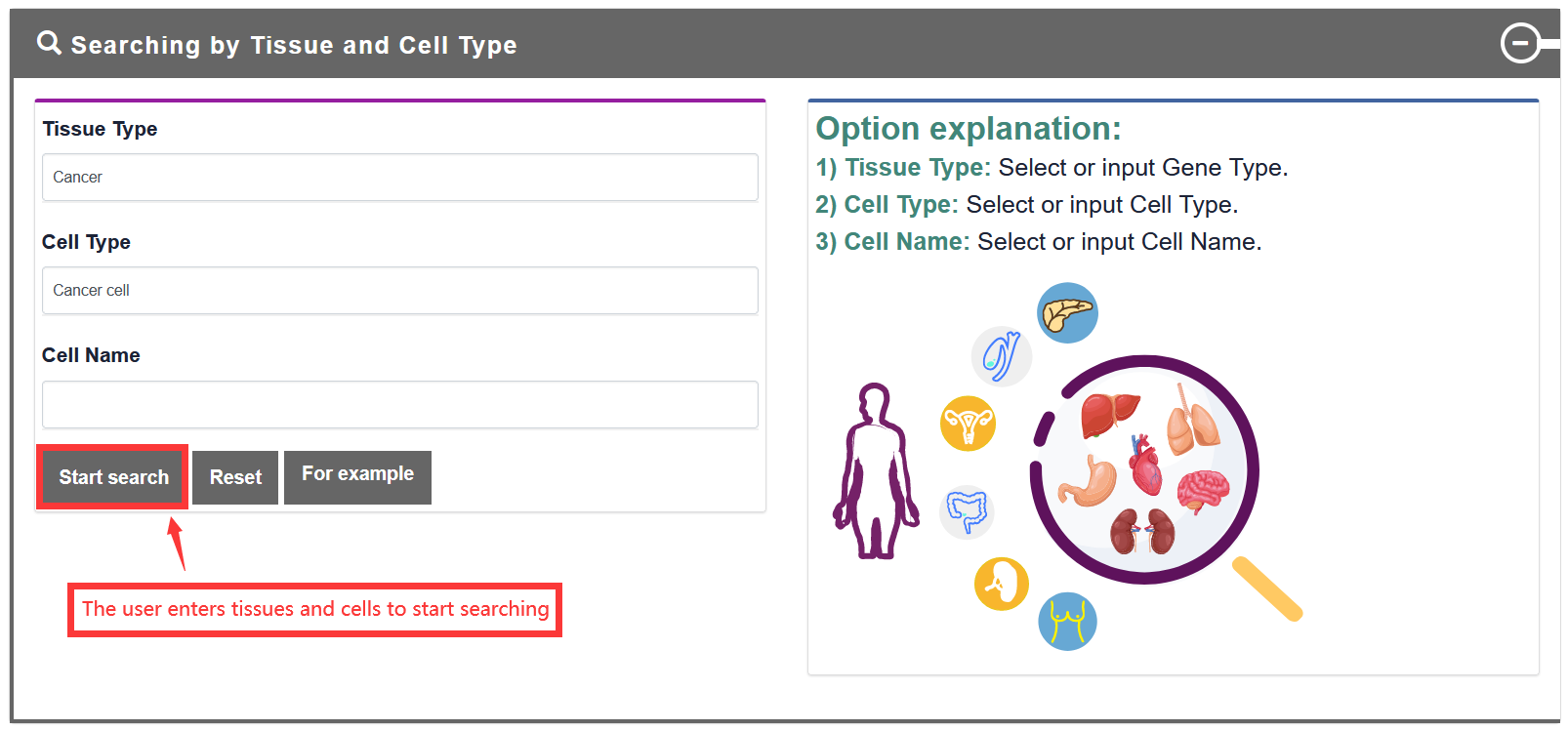
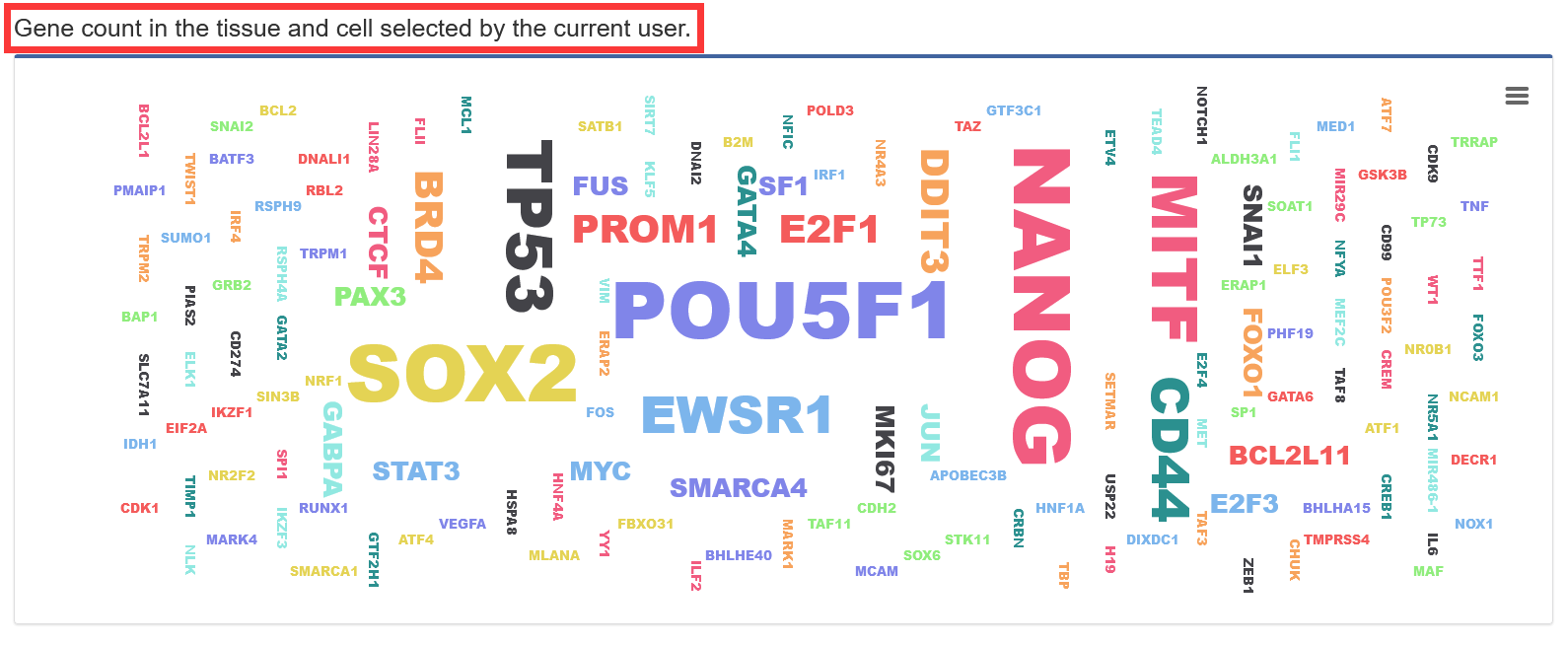
Users searches for tissues and cell types, and TF-Marker will return detailed information, including gene information in tissues and cell types, and text cloud pictures about gene distribution in tissues and cell types.

3.1.2. Search by gene name
Users search for gene name, and TF-Marker will return detailed information, including gene distribution in tissues and cell types, and gene expression in 'GTEx', 'CCLE', 'TCGA' and 'Encode Cell Lines'.
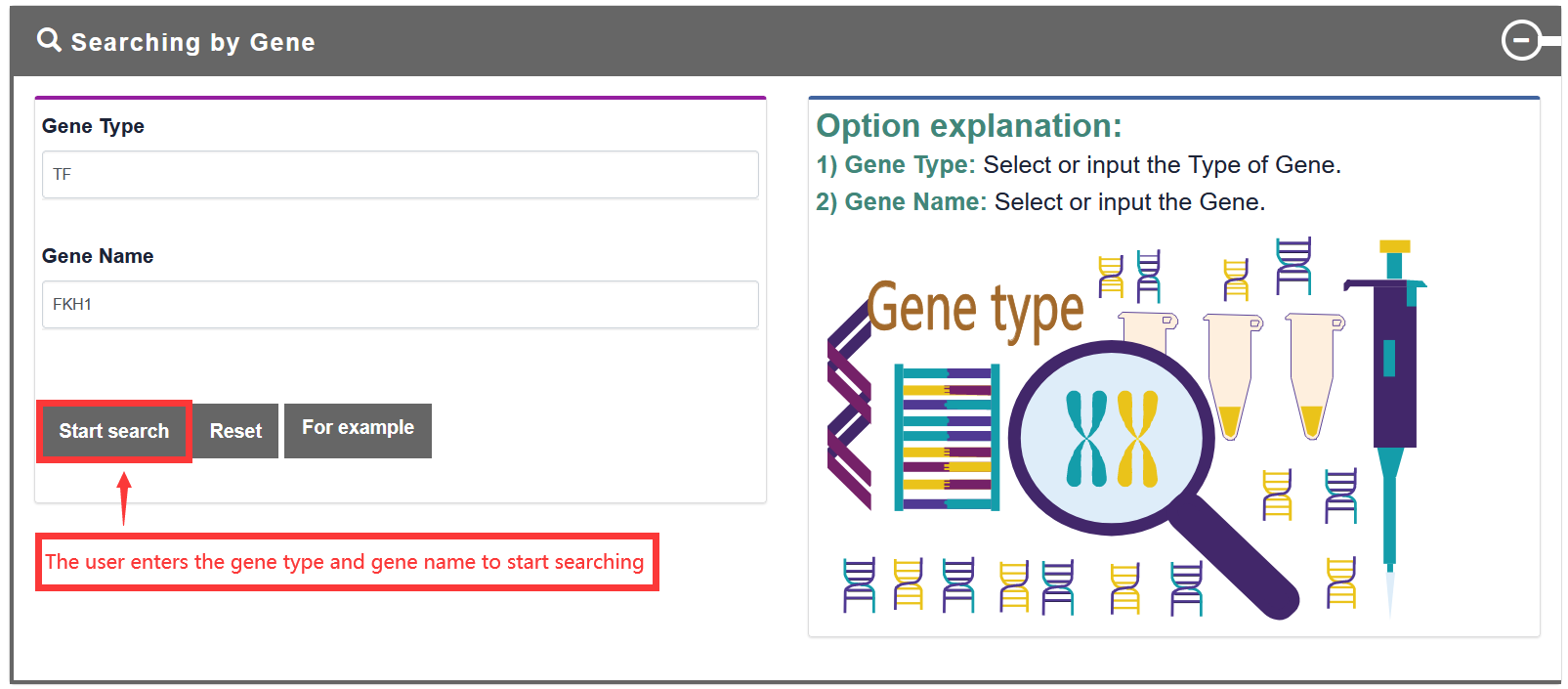
TF-Marker will be automatically converted into a gene symbol if the user inputs the Ensembl ID/NCBI Refseq ID/Alias/Entrez Gene ID
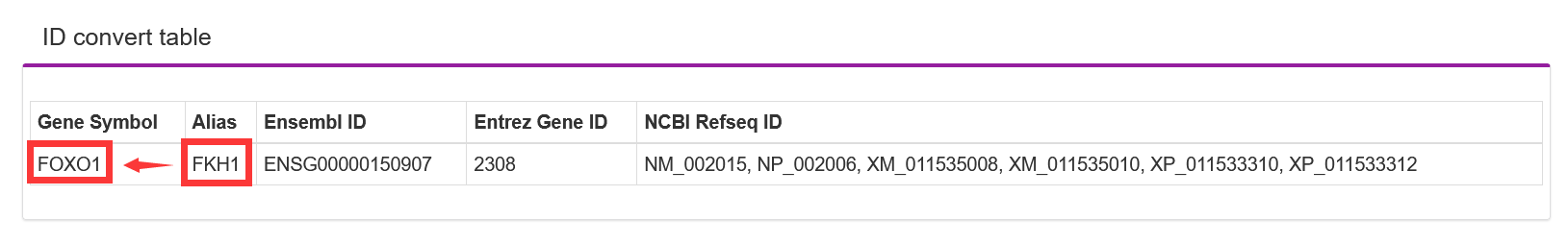
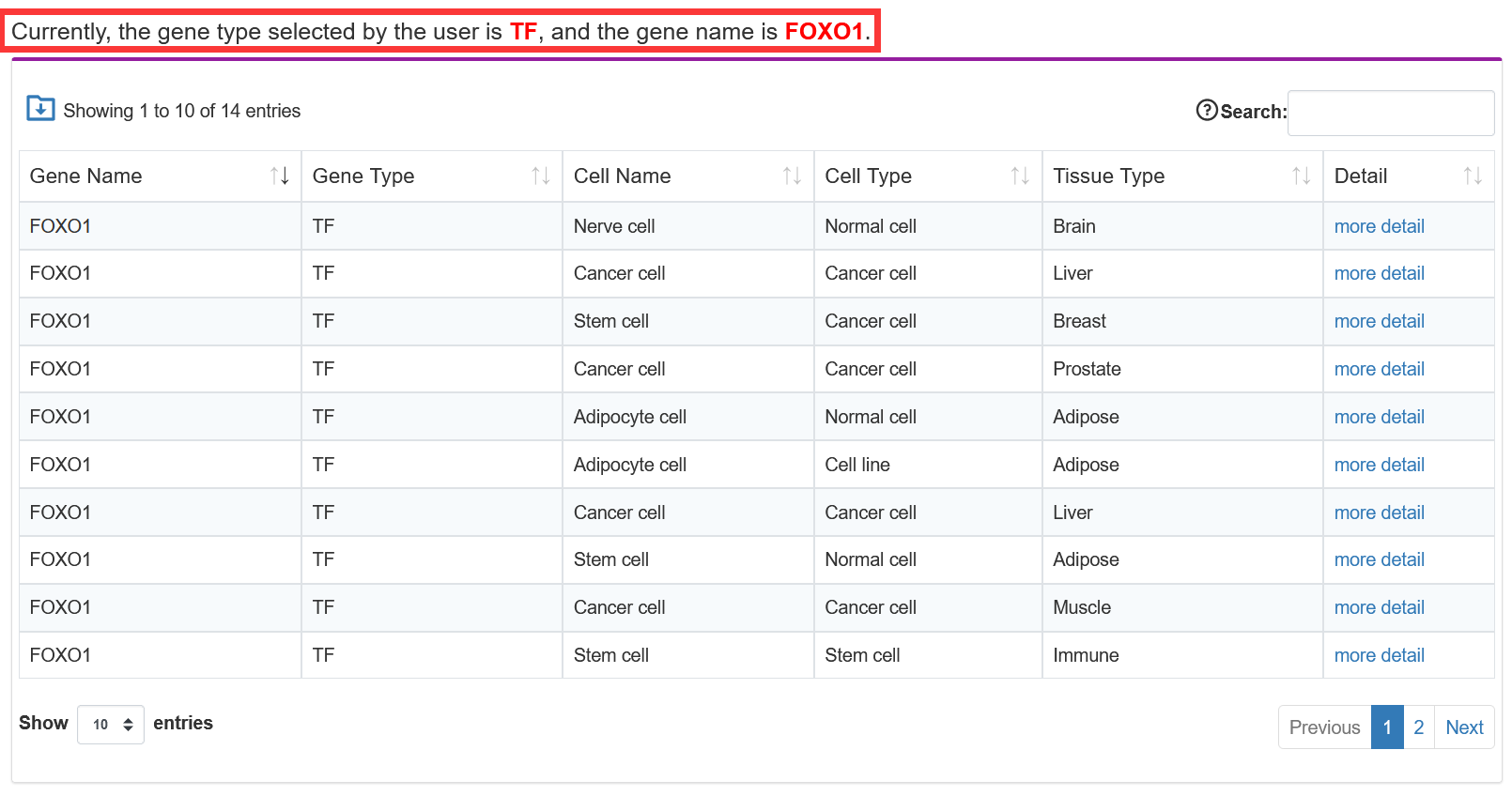


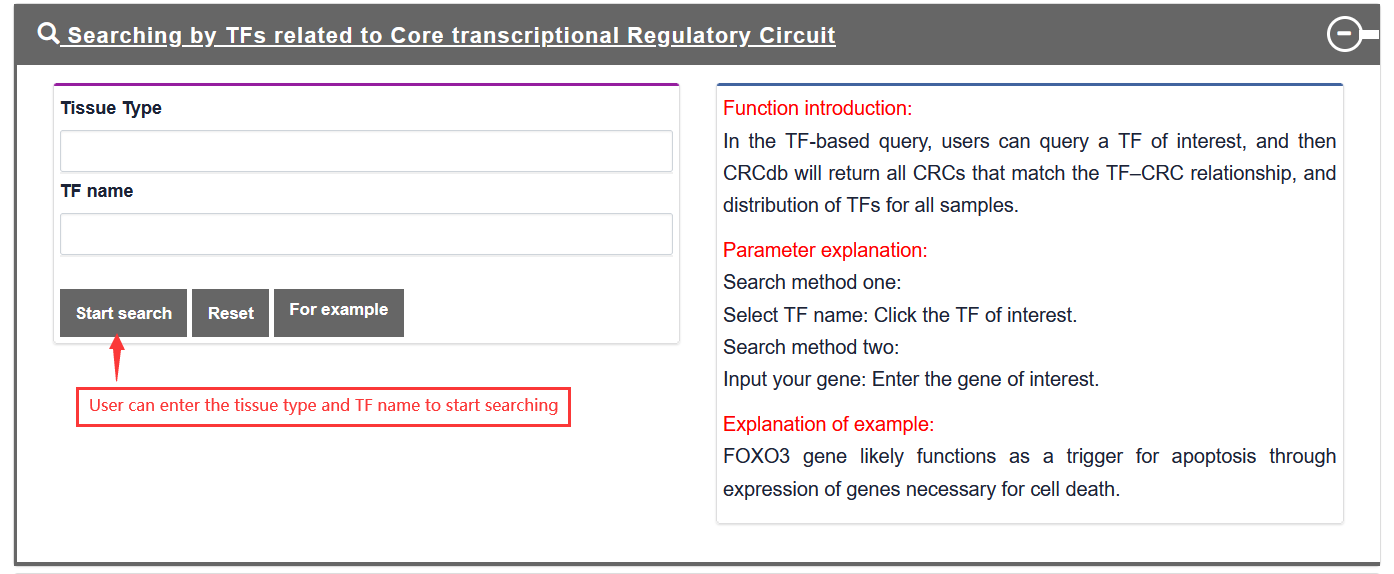
3.1.3. Search by TF related to Core transcriptional Regulatory Circuit
Users search for TF related to Core transcriptional Regulatory Circuit, and TF-Marker will return the TF information about CRC, including TF distribution in the tissue entered by user, and others related to TF downstream and upstream.

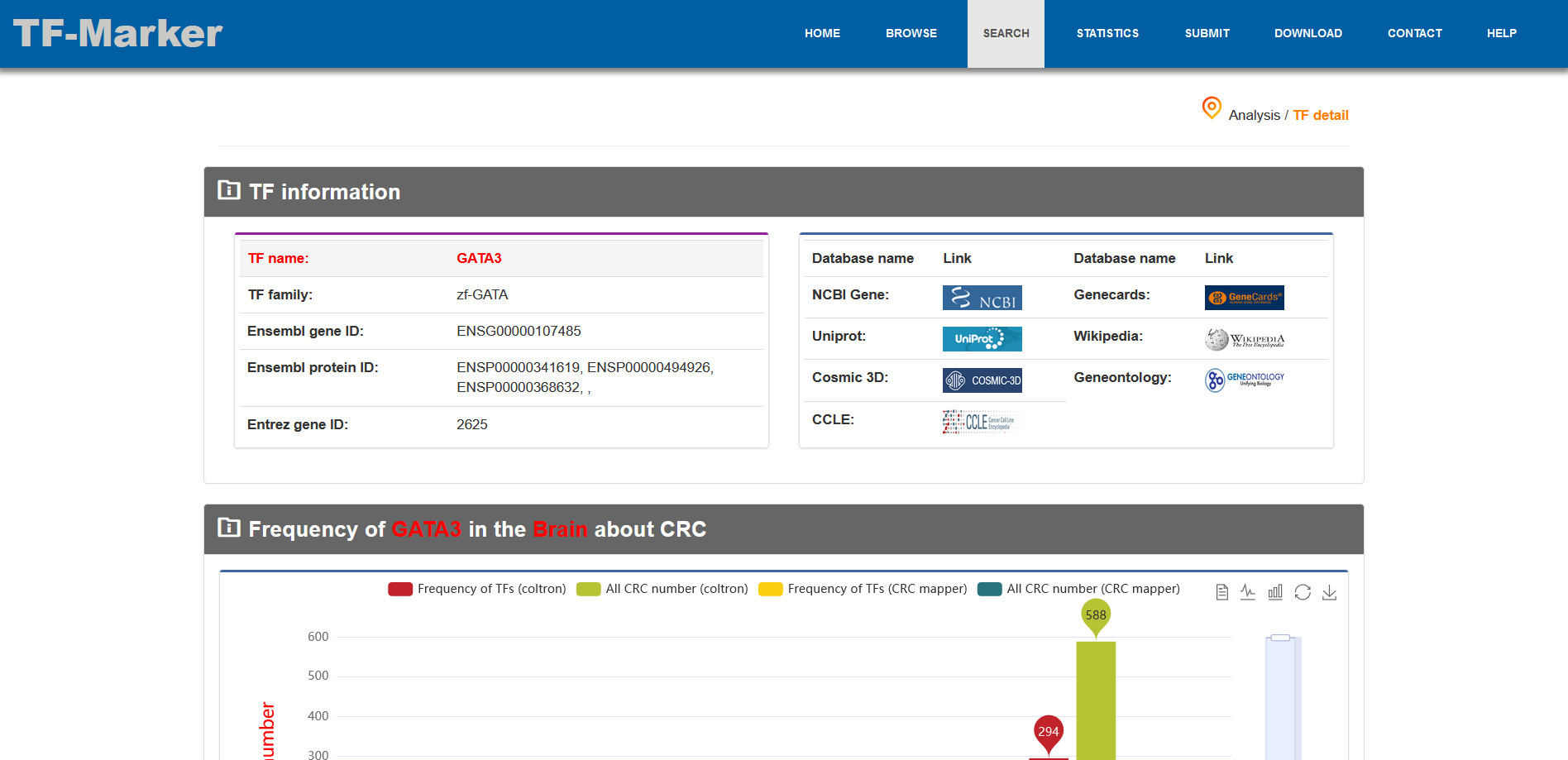


3.2. Browse
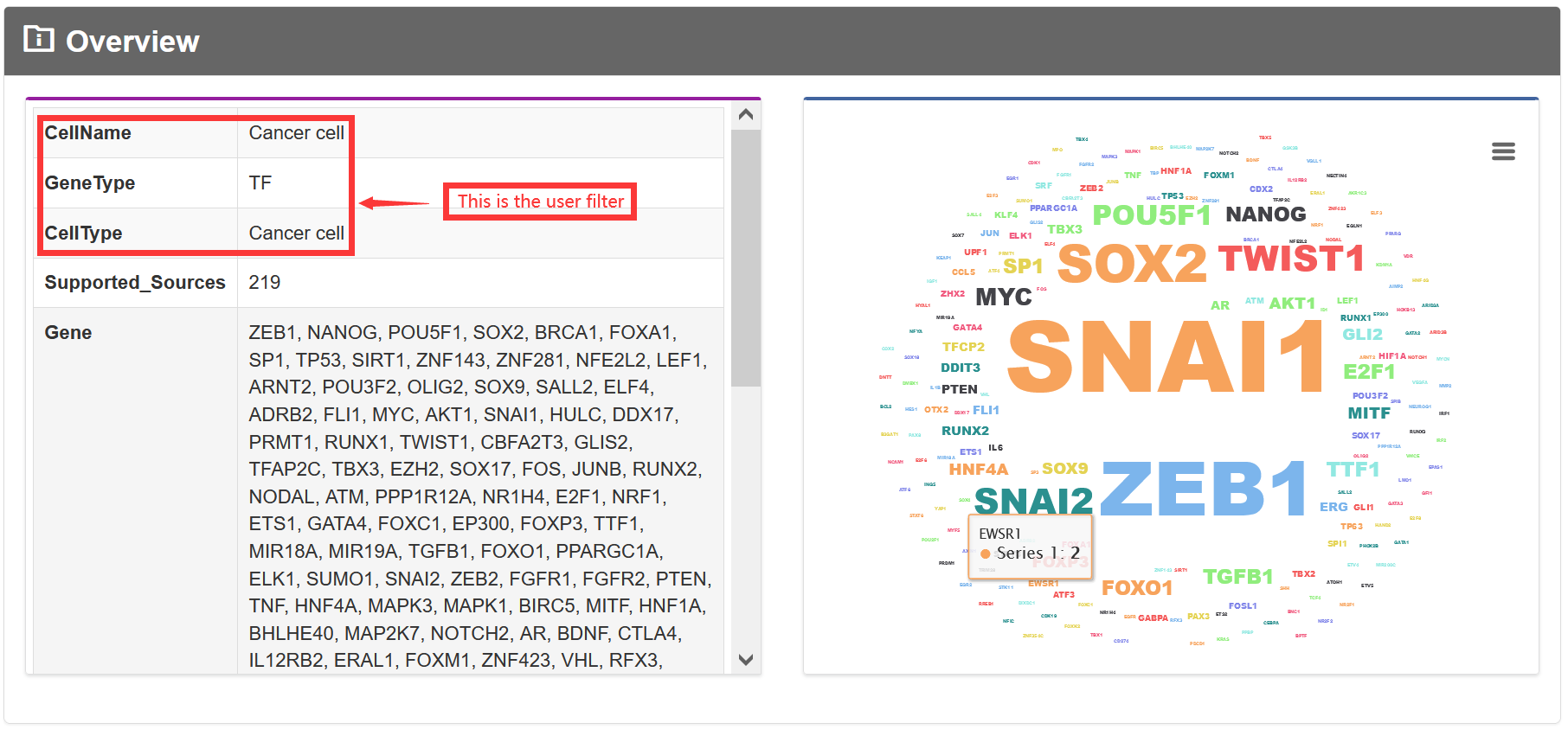
The 'Data-Browse' page is organized as an interactive and alphanumerically sortable table that allows users to quickly browse through 'Tissue type', 'Cell type' and 'Gene Type'. The 'Show entries' drop-down menu is provided for changing the number of records per page conveniently.
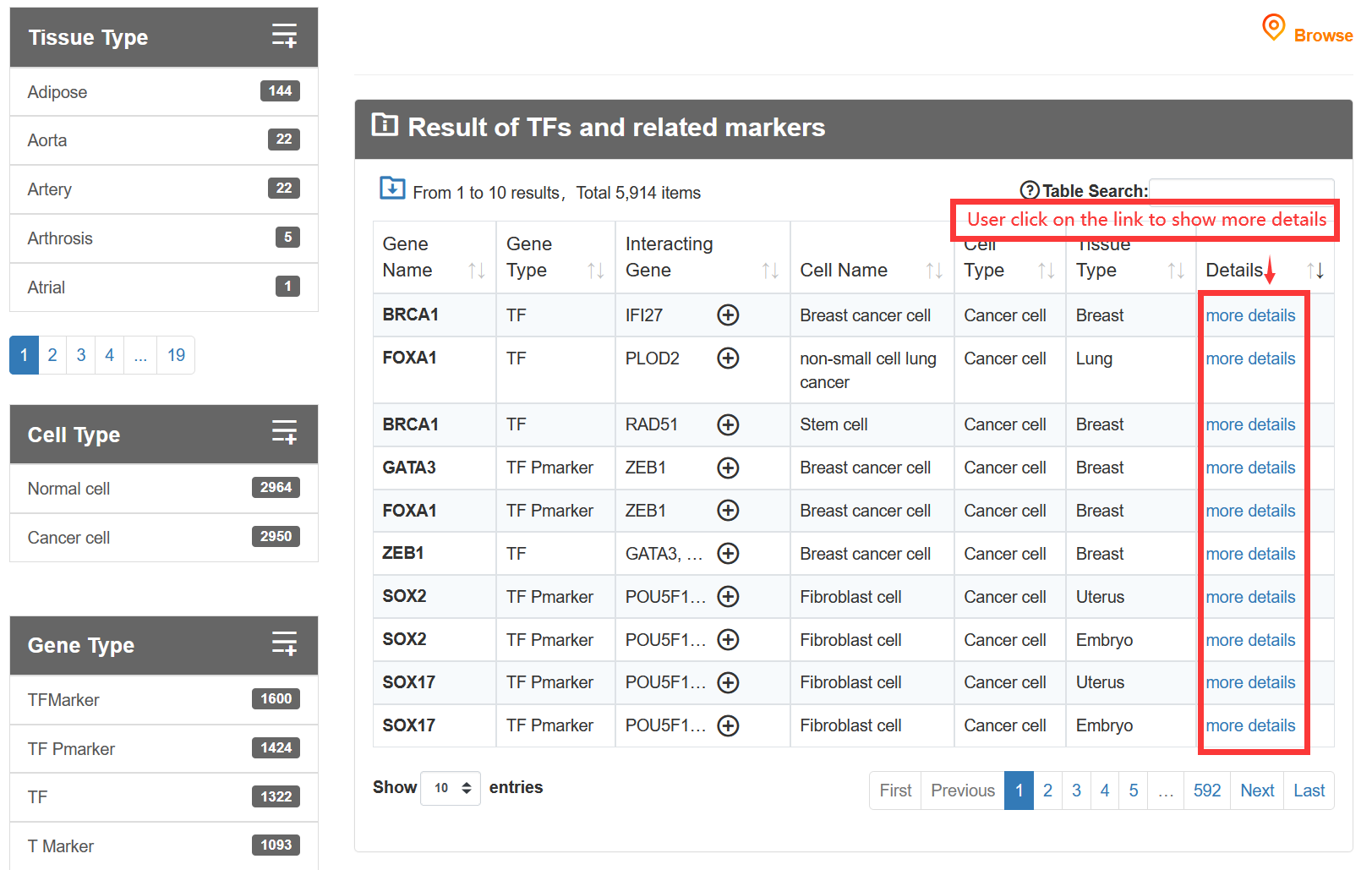


If the gene type is a TF, the graph will be displayed below.
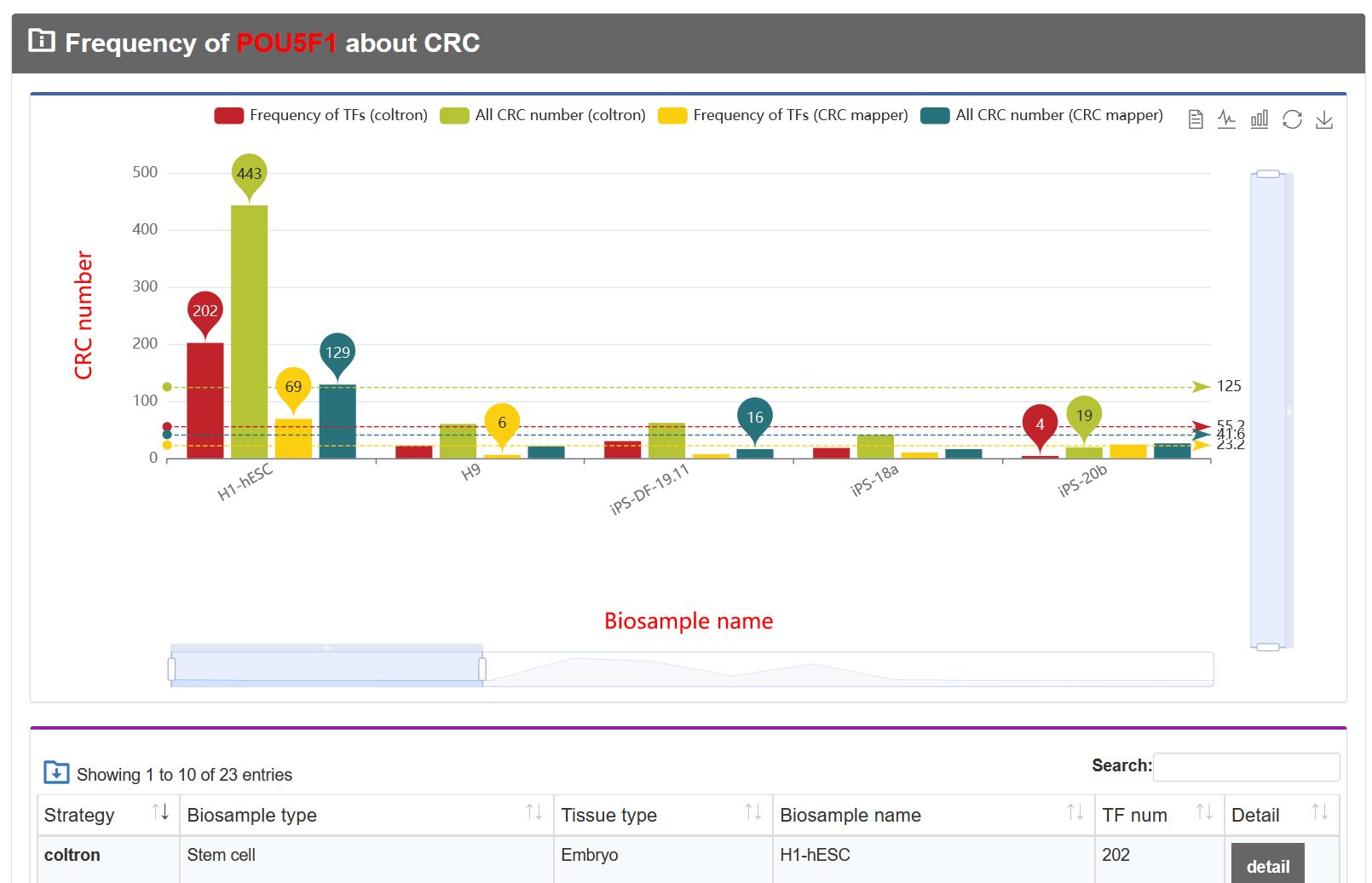
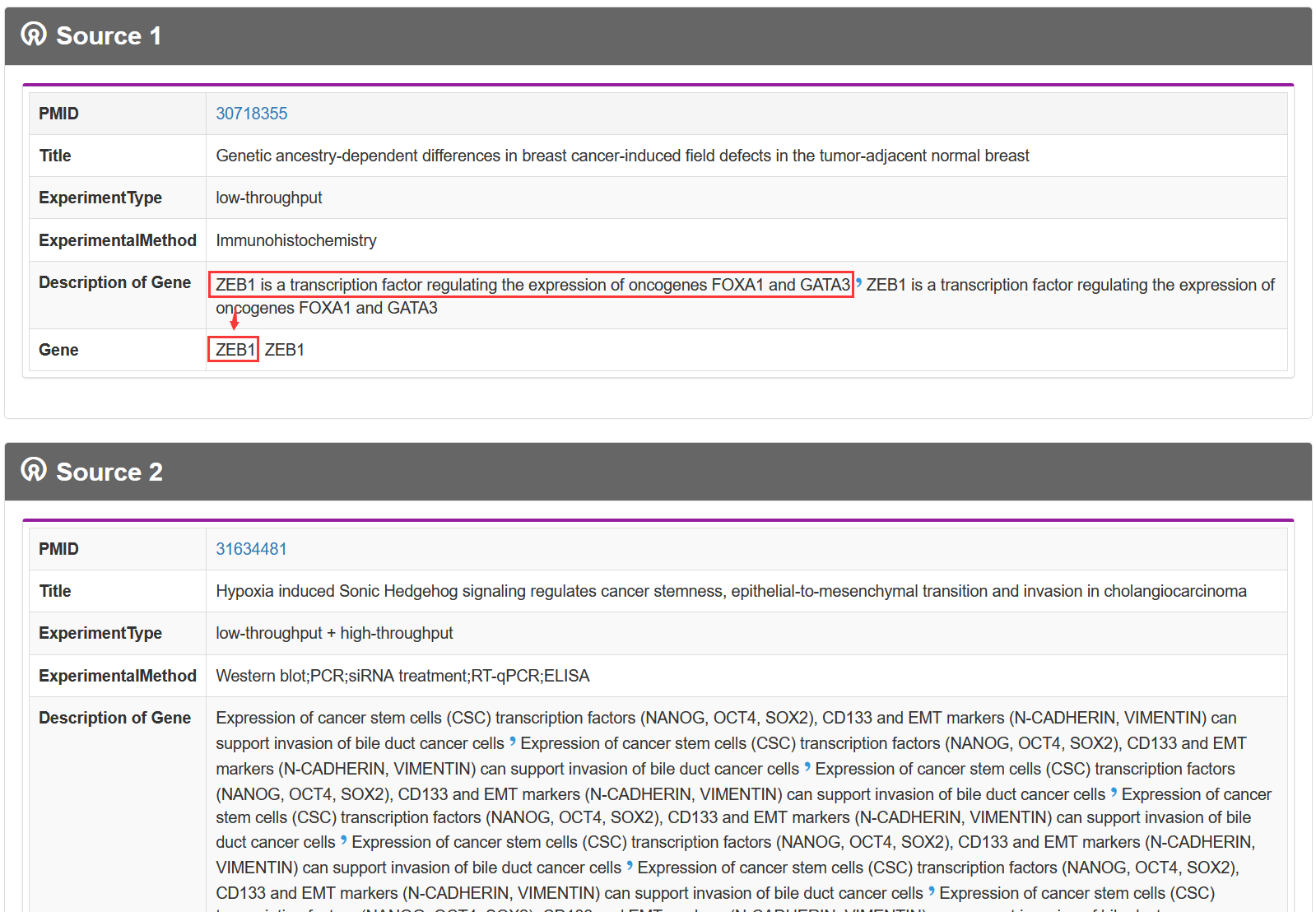
On the 'Result of cell types', TF-Marker will provide information about the genes in cells under the screening conditions of users, and the details page will show the specific sources of genes and some experimental information of corresponding articles.
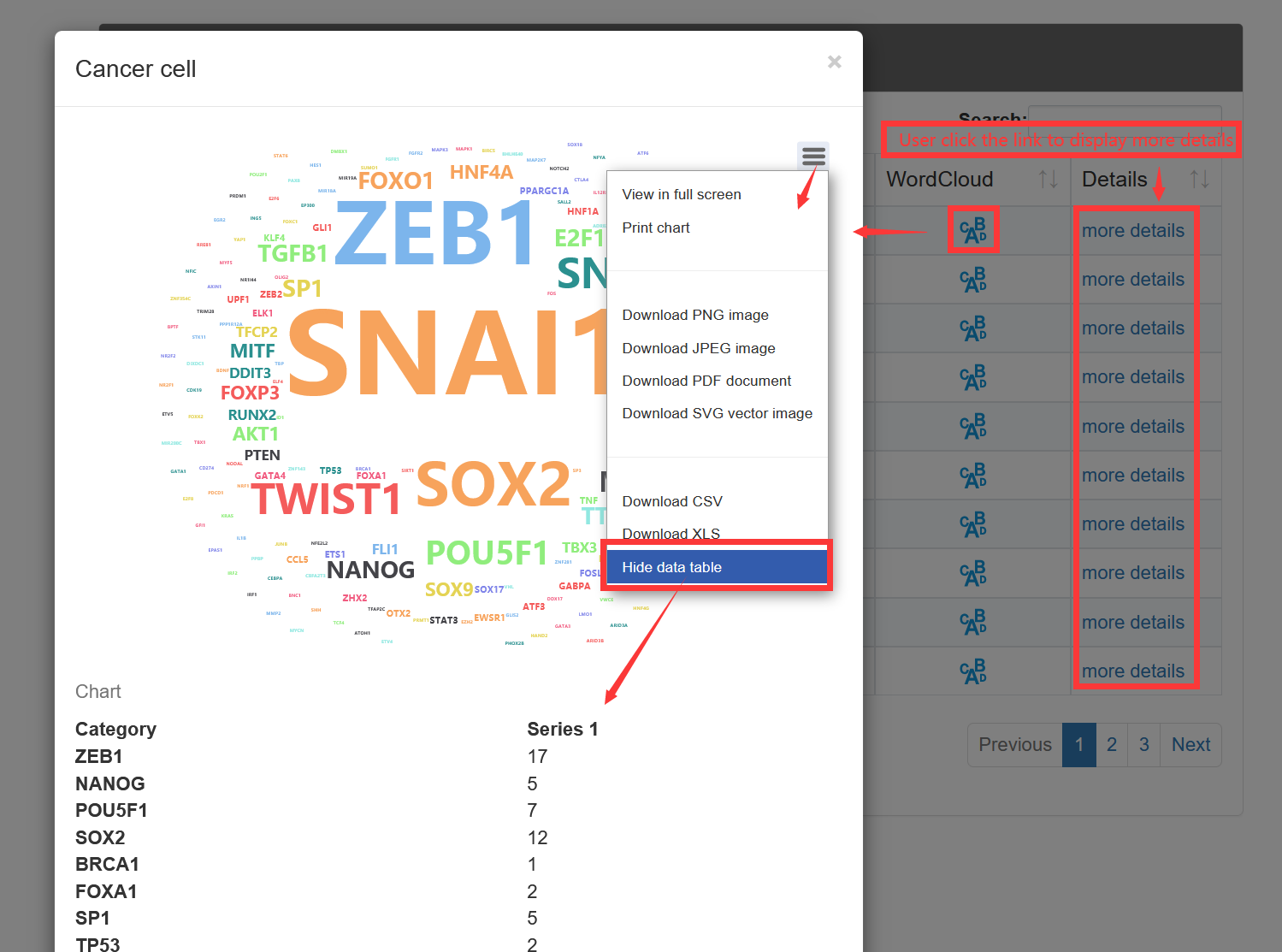
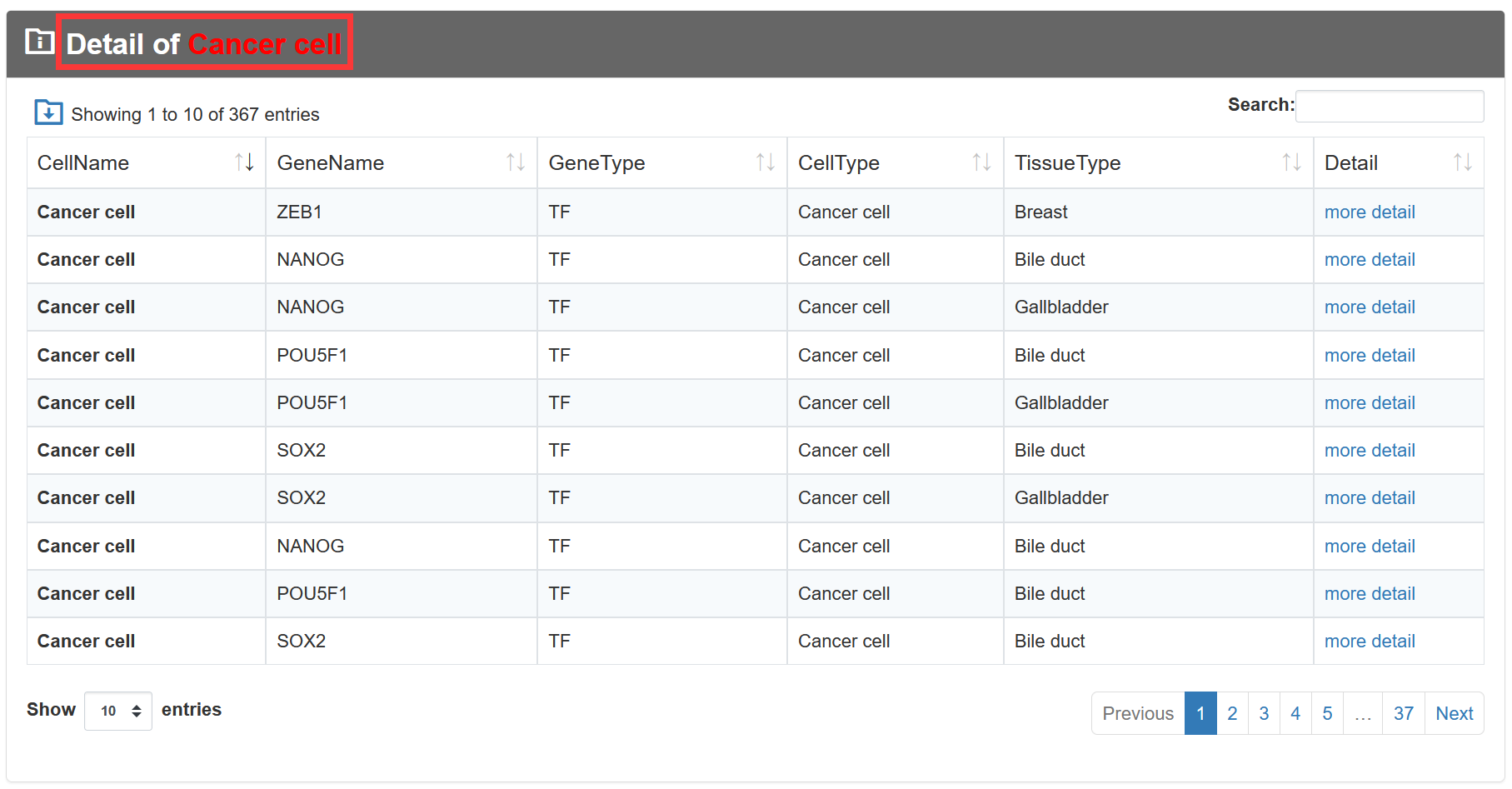
3.3. Submit
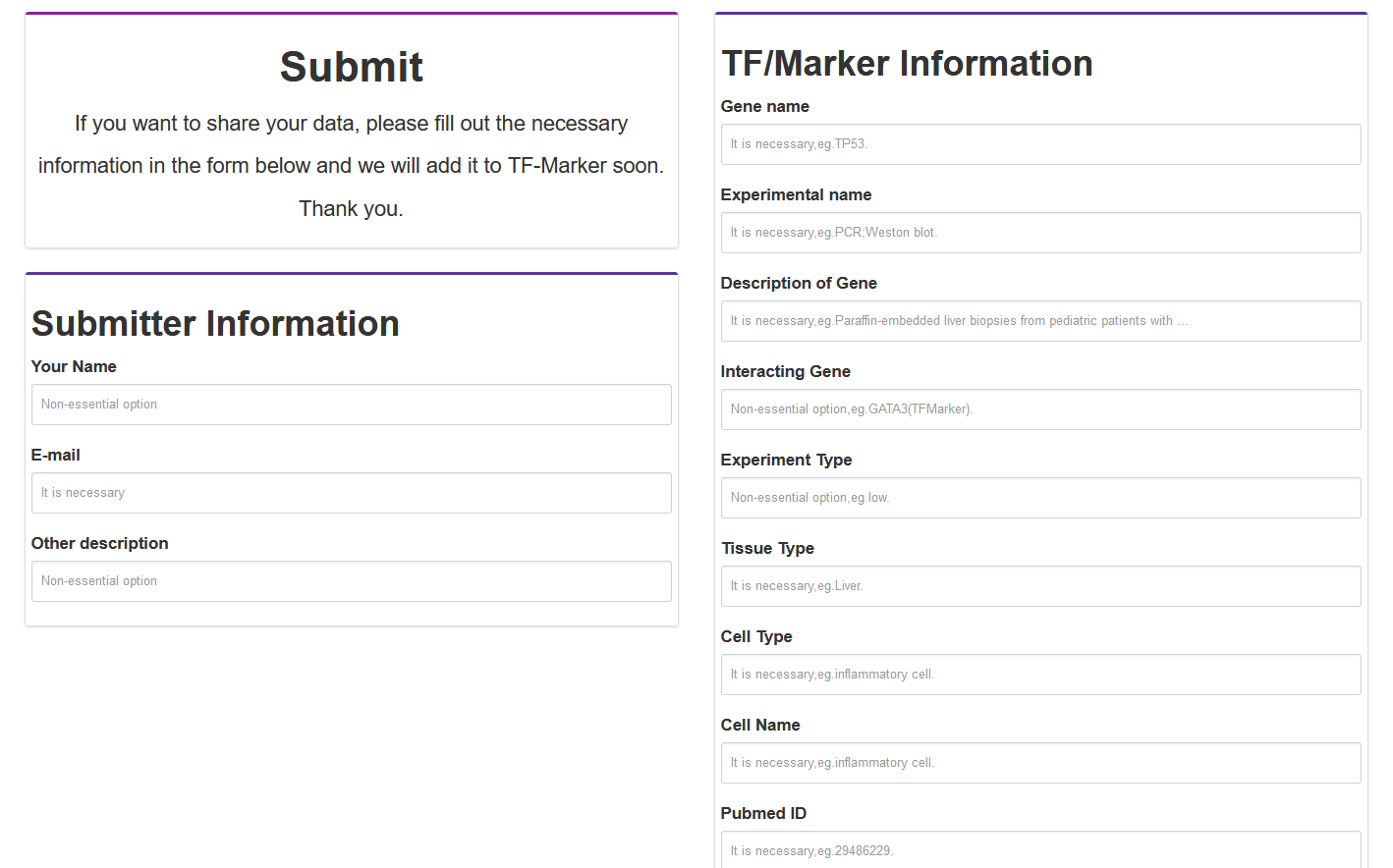
Users can submit 'Gene Name', 'Gene Type', 'Cell Name', 'Gene ID', 'Ensembl ID', 'Transcription factor Family', 'Tissue type', 'experiment name', 'Pubmed ID' and the description of TFs and related markers in the literature through the submit page.
3.4 Download
The 'Download' page exhibits 'Gene Name', 'Gene Type', 'Cell Name', 'Gene ID', 'Ensembl ID', 'Transcription factor Family', 'Tissue type', 'experiment name', 'Pubmed ID' and the description of TFs and related markers in the literature for users to download. Moreover, the detailed description of file is also displayed.
4. Explanation of the definitions used by the website
| Attribution | The description of the attribution |
|---|---|
| Gene Name | The named gene in the article that can be homologous to human |
| Gene Symbol | The name of the gene in GENE database that can be homologous to human. |
| Gene Type | According to the description of genes,the genes can be divided into 5 categories. |
| Gene ID | The information of the gene in GENE database. |
| Ensembl ID | The information of the gene in Ensembl database. |
| Transcription factor Family | Transcription factor family information to descripe the TF. |
| Interacting Gene | Interacting genes are described in the paper. |
| Cell Name | The names of the cells studied in this paper. |
| Cell Type | The types of the cells studied in this paper. |
| Tissue Type | The types of the tissues studied in this paper. |
| Experiment Type | Experimental categories( High throughput/Low throughput). |
| Experiment Name | Experiments used to study the gene. |
| PMID | PMID of the article in PubMed. |
| Title | The title of the article. |
| Description | The description of the gene in the article. |
4. Development environment
Using MySQL 5.7.27, we developed the current version of TF-Marker that runs on a Linux-based Apache web server. We utilized PHP 5.6.40 for sever-side scripting, Bootstrap v3.37 and JQuerry v2.1.1 for interactive interface building, Echats for visualization. To display best, we recommend using a comprehensive web sever that supports HTML5 standard, for example, Firefox, Google Chrome and Safari.
The research community can access information freely in TF-Marker database without registering or logging in. The web link of TF-Marker is http://www.licpathway.net/TF-Marker.